mTOR
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 1: 11.11 – 11.26 Mb | Chr 4: 148.53 – 148.64 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
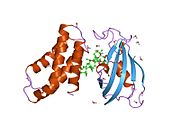
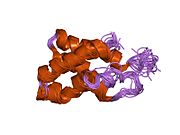
The mammalian target of rapamycin (mTOR),[5] also referred to as the mechanistic target of rapamycin, and sometimes called FK506-binding protein 12-rapamycin-associated protein 1 (FRAP1), is a kinase that in humans is encoded by the MTOR gene.[6][7][8] mTOR is a member of the phosphatidylinositol 3-kinase-related kinase family of protein kinases.[9]
mTOR links with other proteins and serves as a core component of two distinct
Discovery
Rapa Nui (Easter Island - Chile)
The study of TOR originated in the 1960s with an expedition to
Subsequent history
The discovery of TOR and mTOR stemmed from independent studies of the natural product rapamycin by Joseph Heitman, Rao Movva, and Michael N. Hall in 1991;[16] by David M. Sabatini, Hediye Erdjument-Bromage, Mary Lui, Paul Tempst, and Solomon H. Snyder[7] in 1994; and by Candace J. Sabers, Mary M. Martin, Gregory J. Brunn, Josie M. Williams, Francis J. Dumont, Gregory Wiederrecht, and Robert T. Abraham in 1995.[8] In 1991, working in yeast, Hall and colleagues identified the TOR1 and TOR2 genes.[16] In 1993, Robert Cafferkey, George Livi, and colleagues, and Jeannette Kunz, Michael N. Hall, and colleagues independently cloned genes that mediate the toxicity of rapamycin in fungi, known as the TOR/DRR genes.[17][18] However, the molecular target of the FKBP12-rapamycin complex in mammals was not known. In 1994, researchers working in the labs of Stuart L. Schreiber, Solomon H. Snyder and Robert T. Abraham independently discovered a protein that directly interacts with FKBP12-rapamycin, which became known as mTOR due to its homology to the yeast TOR/DRR genes.[6][7][8]
Rapamycin arrests fungal activity at the
In 1991, calcineurin was identified as the target of FKBP12-FK506.[27] That of FKBP12-rapamycin remained mysterious until genetic and molecular studies in yeast established FKBP12 as the target of rapamycin, and implicated TOR1 and TOR2 as the targets of FKBP12-rapamycin in 1991 and 1993,[16][28] followed by studies in 1994 when several groups, working independently, discovered the mTOR kinase as its direct target in mammalian tissues.[6][7][20] Sequence analysis of mTOR revealed it to be the direct ortholog of proteins encoded by the yeast target of rapamycin 1 and 2 (TOR1 and TOR2) genes, which Joseph Heitman, Rao Movva, and Michael N. Hall had identified in August 1991 and May 1993. Independently, George Livi and colleagues later reported the same genes, which they called dominant rapamycin resistance 1 and 2 (DRR1 and DRR2), in studies published in October 1993.
The protein, now called mTOR, was originally named FRAP by Stuart L. Schreiber and RAFT1 by David M. Sabatini;[6][7] FRAP1 was used as its official gene symbol in humans. Because of these different names, mTOR, which had been first used by Robert T. Abraham,[6] was increasingly adopted by the community of scientists working on the mTOR pathway to refer to the protein and in homage to the original discovery of the TOR protein in yeast that was named TOR, the Target of Rapamycin, by Joe Heitman, Rao Movva, and Mike Hall. TOR was originally discovered at the Biozentrum and Sandoz Pharmaceuticals in 1991 in Basel, Switzerland, and the name TOR pays further homage to this discovery, as TOR means doorway or gate in German, and the city of Basel was once ringed by a wall punctuated with gates into the city, including the iconic Spalentor.[29] "mTOR" initially meant "mammalian target of rapamycin", but the meaning of the "m" was later changed to "mechanistic".[30] Similarly, with subsequent discoveries the zebra fish TOR was named zTOR, the Arabidopsis thaliana TOR was named AtTOR, and the Drosophila TOR was named dTOR. In 2009 the FRAP1 gene name was officially changed by the HUGO Gene Nomenclature Committee (HGNC) to mTOR, which stands for mechanistic target of rapamycin.[31]
The discovery of TOR and the subsequent identification of mTOR opened the door to the molecular and physiological study of what is now called the mTOR pathway and had a catalytic effect on the growth of the field of chemical biology, where small molecules are used as probes of biology.
Function
mTOR integrates the input from upstream
In plants
Plants express the mechanistic target of rapamycin (mTOR) and have a TOR kinase complex. In plants, only the TORC1 complex is present unlike that of mammalian target of rapamycin which also contains the TORC2 complex.[38] Plant species have TOR proteins in the protein kinase and FKBP-rapamycin binding (FRB) domains that share a similar amino acid sequence to mTOR in mammals.[39]
Role of mTOR in plants
The TOR kinase complex has been known for having a role in the metabolism of plants. The TORC1 complex turns on when plants are living the proper environmental conditions to survive. Once activated, plant cells undergo particular anabolic reactions. These include plant development, translation of mRNA and the growth of cells within the plant. However, the TORC1 complex activation stops catabolic processes such as autophagy from occurring.[38] TOR kinase signaling in plants has been found to aid in senescence, flowering, root and leaf growth, embryogenesis, and the meristem activation above the root cap of a plant. [40] mTOR is also found to be highly involved in developing embryo tissue in plants.[39]
Complexes
mTOR is the
mTORC1
mTOR Complex 1 (mTORC1) is composed of mTOR, regulatory-associated protein of mTOR (
mTORC2
mTOR Complex 2 (mTORC2) is composed of MTOR, rapamycin-insensitive companion of MTOR (
Inhibition by rapamycin
Rapamycin (Sirolimus) inhibits mTORC1, resulting in the suppression of cellular senescence.[54] This appears to provide most of the beneficial effects of the drug (including life-span extension in animal studies). Suppression of insulin resistance by sirtuins accounts for at least some of this effect.[55] Impaired sirtuin 3 leads to mitochondrial dysfunction.[56]
Rapamycin has a more complex effect on mTORC2, inhibiting it only in certain cell types under prolonged exposure. Disruption of mTORC2 produces the diabetic-like symptoms of decreased glucose tolerance and insensitivity to insulin.[57]
Gene deletion experiments
The mTORC2 signaling pathway is less defined than the mTORC1 signaling pathway. The functions of the components of the mTORC complexes have been studied using knockdowns and knockouts and were found to produce the following phenotypes:
- NIP7: Knockdown reduced mTORC2 activity that is indicated by decreased phosphorylation of mTORC2 substrates.[58]
- RICTOR: Overexpression leads to metastasis and knockdown inhibits growth factor-induced PKC-phosphorylation.[59] Constitutive deletion of Rictor in mice leads to embryonic lethality,[60] while tissue specific deletion leads to a variety of phenotypes; a common phenotype of Rictor deletion in liver, white adipose tissue, and pancreatic beta cells is systemic glucose intolerance and insulin resistance in one or more tissues.[57][61][62][63] Decreased Rictor expression in mice decreases male, but not female, lifespan.[64]
- mTOR: Inhibition of mTORC1 and mTORC2 by PP242 [2-(4-Amino-1-isopropyl-1H-pyrazolo[3,4-d]pyrimidin-3-yl)-1H-indol-5-ol] leads to autophagy or apoptosis; inhibition of mTORC2 alone by PP242 prevents phosphorylation of Ser-473 site on AKT and arrests the cells in G1 phase of the cell cycle.[65] Genetic reduction of mTOR expression in mice significantly increases lifespan.[66]
- PDK1: Knockout is lethal; hypomorphic allele results in smaller organ volume and organism size but normal AKT activation.[67]
- TOR1, the S. cerevisiae orthologue of mTORC1, is a regulator of both carbon and nitrogen metabolism; TOR1 KO strains regulate response to nitrogen as well as carbon availability, indicating that it is a key nutritional transducer in yeast.[70][71]
Clinical significance
Aging

Decreased TOR activity has been found to increase life span in
It is hypothesized that some dietary regimes, like
According to the
mTOR is a key initiator of the
Cancer
Over-activation of mTOR signaling significantly contributes to the initiation and development of tumors and mTOR activity was found to be deregulated in many types of cancer including breast, prostate, lung, melanoma, bladder, brain, and renal carcinomas.
Increasing mTOR activity was shown to drive cell cycle progression and increase cell proliferation mainly due to its effect on protein synthesis. Moreover, active mTOR supports tumor growth also indirectly by inhibiting
Central nervous system disorders / Brain function
Autism
mTOR is implicated in the failure of a 'pruning' mechanism of the excitatory synapses in autism spectrum disorders.[98]
Alzheimer's disease
mTOR signaling intersects with Alzheimer's disease (AD) pathology in several aspects, suggesting its potential role as a contributor to disease progression. In general, findings demonstrate mTOR signaling hyperactivity in AD brains. For example, postmortem studies of human AD brain reveal dysregulation in PTEN, Akt, S6K, and mTOR.[99][100][101] mTOR signaling appears to be closely related to the presence of soluble amyloid beta (Aβ) and tau proteins, which aggregate and form two hallmarks of the disease, Aβ plaques and neurofibrillary tangles, respectively.[102] In vitro studies have shown Aβ to be an activator of the PI3K/AKT pathway, which in turn activates mTOR.[103] In addition, applying Aβ to N2K cells increases the expression of p70S6K, a downstream target of mTOR known to have higher expression in neurons that eventually develop neurofibrillary tangles.[104][105] Chinese hamster ovary cells transfected with the 7PA2 familial AD mutation also exhibit increased mTOR activity compared to controls, and the hyperactivity is blocked using a gamma-secretase inhibitor.[106][107] These in vitro studies suggest that increasing Aβ concentrations increases mTOR signaling; however, significantly large, cytotoxic Aβ concentrations are thought to decrease mTOR signaling.[108]
Consistent with data observed in vitro, mTOR activity and activated p70S6K have been shown to be significantly increased in the cortex and hippocampus of animal models of AD compared to controls.[107][109] Pharmacologic or genetic removal of the Aβ in animal models of AD eliminates the disruption in normal mTOR activity, pointing to the direct involvement of Aβ in mTOR signaling.[109] In addition, by injecting Aβ oligomers into the hippocampi of normal mice, mTOR hyperactivity is observed.[109] Cognitive impairments characteristic of AD appear to be mediated by the phosphorylation of PRAS-40, which detaches from and allows for the mTOR hyperactivity when it is phosphorylated; inhibiting PRAS-40 phosphorylation prevents Aβ-induced mTOR hyperactivity.[109][110][111] Given these findings, the mTOR signaling pathway appears to be one mechanism of Aβ-induced toxicity in AD.
The hyperphosphorylation of tau proteins into neurofibrillary tangles is one hallmark of AD. p70S6K activation has been shown to promote tangle formation as well as mTOR hyperactivity through increased phosphorylation and reduced dephosphorylation.[104][112][113][114] It has also been proposed that mTOR contributes to tau pathology by increasing the translation of tau and other proteins.[115]
Synaptic plasticity is a key contributor to learning and memory, two processes that are severely impaired in AD patients. Translational control, or the maintenance of protein homeostasis, has been shown to be essential for neural plasticity and is regulated by mTOR.[107][116][117][118][119] Both protein over- and under-production via mTOR activity seem to contribute to impaired learning and memory. Furthermore, given that deficits resulting from mTOR overactivity can be alleviated through treatment with rapamycin, it is possible that mTOR plays an important role in affecting cognitive functioning through synaptic plasticity.[103][120] Further evidence for mTOR activity in neurodegeneration comes from recent findings demonstrating that eIF2α-P, an upstream target of the mTOR pathway, mediates cell death in prion diseases through sustained translational inhibition.[121]
Some evidence points to mTOR's role in reduced Aβ clearance as well. mTOR is a negative regulator of autophagy;[122] therefore, hyperactivity in mTOR signaling should reduce Aβ clearance in the AD brain. Disruptions in autophagy may be a potential source of pathogenesis in protein misfolding diseases, including AD.[123][124][125][126][127][128] Studies using mouse models of Huntington's disease demonstrate that treatment with rapamycin facilitates the clearance of huntingtin aggregates.[129][130] Perhaps the same treatment may be useful in clearing Aβ deposits as well.
Lymphoproliferative diseases
Hyperactive mTOR pathways have been identified in certain lymphoproliferative diseases such as autoimmune lymphoproliferative syndrome (ALPS),[131] multicentric Castleman disease,[132] and post-transplant lymphoproliferative disorder (PTLD).[133]
Protein synthesis and cell growth
mTORC1 activation is required for myofibrillar muscle protein synthesis and skeletal
Lysosomal damage inhibits mTOR and induces autophagy
Active
Additionally, several types of ubiquitination events parallel and complement the galectin-driven processes:
Scleroderma
mTOR inhibitors as therapies
Transplantation
mTOR inhibitors, e.g.
Glycogen storage disease
Some articles reported that rapamycin can inhibit mTORC1 so that the phosphorylation of GS (glycogen synthase) can be increased in skeletal muscle. This discovery represents a potential novel therapeutic approach for glycogen storage disease that involve glycogen accumulation in muscle.
Anti-cancer
There are two primary mTOR inhibitors used in the treatment of human cancers, temsirolimus and everolimus. mTOR inhibitors have found use in the treatment of a variety of malignancies, including renal cell carcinoma (temsirolimus) and pancreatic cancer, breast cancer, and renal cell carcinoma (everolimus).[161] The complete mechanism of these agents is not clear, but they are thought to function by impairing tumour angiogenesis and causing impairment of the G1/S transition.[162]
Anti-aging
mTOR inhibitors may be useful for treating/preventing several age-associated conditions,[163] including neurodegenerative diseases such as Alzheimer's disease and Parkinson's disease.[164] After a short-term treatment with the mTOR inhibitors dactolisib and everolimus, in elderly (65 and older), treated subjects had a reduced number of infections over the course of a year.[165]
Various natural compounds, including
Interactions
Mechanistic target of rapamycin has been shown to
- ABL1,[172]
- AKT1,[52][173][174]
- IGF-IR,[12]
- InsR,[12]
- CLIP1,[175]
- EIF3F[176]
- EIF4EBP1,[45][177][178][179][180][181][182][183]
- FKBP1A,[13][50][184][185][186][187]
- GPHN,[188]
- PRKCD,[201]
- RHEB,[180][202][203][204]
- RICTOR,[13][49][50][191][197][199][200]
- RPS6KB1,[45][178][180][181][182][196][199][205][206][207][208][209][210][211][212]
- STAT1,[213]
- STAT3,[214][215]
- and
- UBQLN1.[217]
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000198793 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000028991 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- PMID 7822316.
- ^ S2CID 4359651.
- ^ S2CID 33647539.
- ^ PMID 7822316.
- ^ PMID 26134944.
- ^ PMID 25374355.
The mTOR signaling pathway acts as a molecular systems integrator to support organismal and cellular interactions with the environment. The mTOR pathway regulates homeostasis by directly influencing protein synthesis, transcription, autophagy, metabolism, and organelle biogenesis and maintenance. It is not surprising then that mTOR signaling is implicated in the entire hierarchy of brain function including the proliferation of neural stem cells, the assembly and maintenance of circuits, experience-dependent plasticity and regulation of complex behaviors like feeding, sleep and circadian rhythms. ...
mTOR function is mediated through two large biochemical complexes defined by their respective protein composition and have been extensively reviewed elsewhere(Dibble and Manning, 2013; Laplante and Sabatini, 2012)(Figure 1B). In brief, common to both mTOR complex 1 (mTORC1) and mTOR complex 2 (mTORC2) are: mTOR itself, mammalian lethal with sec13 protein 8 (mLST8; also known as GβL), and the inhibitory DEP domain containing mTOR-interacting protein (DEPTOR). Specific to mTORC1 is the regulator-associated protein of the mammalian target of rapamycin (Raptor) and proline-rich Akt substrate of 40 kDa (PRAS40)(Kim et al., 2002; Laplante and Sabatini, 2012). Raptor is essential to mTORC1 activity. The mTORC2 complex includes the rapamycin insensitive companion of mTOR (Rictor), mammalian stress activated MAP kinase-interacting protein 1 (mSIN1), and proteins observed with rictor 1 and 2 (PROTOR 1 and 2)(Jacinto et al., 2006; Jacinto et al., 2004; Pearce et al., 2007; Sarbassov et al., 2004)(Figure 1B). Rictor and mSIN1 are both critical to mTORC2 function.
Figure 1: Domain structure of the mTOR kinase and components of mTORC1 and mTORC2
Figure 2: The mTOR Signaling Pathway - ^ PMID 15314020.
- ^ PMID 26584640.
- ^ S2CID 13831153.
- PMID 36228182.
- PMID 1102509.
- ^ S2CID 9937225.
- S2CID 42926249.
- PMID 8413204.
- ^ S2CID 12932678.
- ^ PMID 8717522.
- PMID 2123553.
- S2CID 11123023.
- S2CID 44256944.
- S2CID 13201695.
- S2CID 4349152.
- .
- S2CID 22094672.
- S2CID 42926249.
- PMID 26540102.
- PMID 27304501.
- ^ "Symbol report for MTOR". HGNC data for MTOR. HUGO Gene Nomenclature Committee. September 1, 2020. Retrieved 2020-12-17.
- PMID 14684182.
- PMID 30803496.
- S2CID 25454463.
- PMID 27304501.
- PMID 12030785.
- ^ PMID 12878853.
- ^ PMID 33138108.
- ^ PMID 29986898.
- PMID 24385567.
- PMID 16469695.
- PMID 24385483.
- S2CID 8237502.
- PMID 22749019.
- ^ PMID 12150925.
- PMID 12718876.
- S2CID 44444716.
- PMID 26937223.
- ^ PMID 16919458.
- ^ PMID 15268862.
- PMID 23852728.
- ^ S2CID 45837814.
- PMID 9445477.
- PMID 34878722.
- PMID 30574122.
- PMID 37329949.
- ^ PMID 22461615.
- PMID 21376236.
- PMID 20978191.
- PMID 17141160.
- PMID 21266327.
- PMID 24072782.
- PMID 20332342.
- PMID 25059582.
- PMID 19209957.
- PMID 23994476.
- PMID 12110585.
- PMID 15046607.
- PMID 23935948.
- PMID 12456783.
- PMID 16407266.
- ^ PMID 16418483.
- ^ S2CID 42188272.
- S2CID 10377667.
- PMID 15186745.
- PMID 19587680.
- PMID 24341993.
- PMID 24409289.
- PMID 27091134.
- PMID 24556924.
- PMID 26695882.
- PMID 26185979.
- S2CID 6526786.
- ^ PMID 20585546.
- ^ PMID 16847060.
- PMID 32397145.
- ^ PMID 26147250.
- PMID 28371119.
- S2CID 4960885.
- PMID 29190625.
- PMID 25450580.
- PMID 16002336.
- PMID 22408430.
- S2CID 19250234.
- PMID 21157483.
- S2CID 1853822.
- PMID 23863156.
- PMID 25155956.
- PMID 18598780.
- S2CID 43085490.
- S2CID 54255848.
- PMID 18359102.
- ^ PMID 22202101.
- ^ PMID 12875979.
- S2CID 22153842.
- PMID 8021238.
- ^ PMID 20178983.
- S2CID 8464608.
- ^ PMID 21266573.
- PMID 17386266.
- PMID 17510057.
- PMID 19210753.
- PMID 17971449.
- PMID 11171037.
- PMID 19648118.
- S2CID 9584832.
- S2CID 2780242.
- PMID 19963289.
- PMID 15016380.
- PMID 18568033.
- PMID 22622579.
- PMID 18670193.
- S2CID 13591350.
- PMID 18930136.
- S2CID 4411895.
- PMID 18266959.
- PMID 18773920.
- PMID 19651785.
- PMID 15146184.
- PMID 19555462.
- ^ Völkl, Simon, et al. "Hyperactive mTOR pathway promotes lymphoproliferation and abnormal differentiation in autoimmune lymphoproliferative syndrome." Blood, The Journal of the American Society of Hematology 128.2 (2016): 227-238. https://doi.org/10.1182/blood-2015-11-685024
- ^ Arenas, Daniel J., et al. "Increased mTOR activation in idiopathic multicentric Castleman disease." Blood 135.19 (2020): 1673-1684. https://doi.org/10.1182/blood.2019002792
- ^ El-Salem, Mouna, et al. "Constitutive activation of mTOR signaling pathway in post-transplant lymphoproliferative disorders." Laboratory Investigation 87.1 (2007): 29-39. https://doi.org/10.1038/labinvest.3700494
- ^ PMID 26010896.
- ^ S2CID 29717535.
- PMID 19150856.
- PMID 26267903.
- S2CID 3953100.
- ^ PMID 24791918.
- ^ PMID 29625033.
- PMID 9461583.
- PMID 15989961.
- ^ PMID 19258318.
- ^ PMID 19225151.
- ^ PMID 19211835.
- PMID 25542097.
- PMID 27728777.
- ^ PMID 27693506.
- PMID 23392225.
- PMID 23262492.
- ^ PMID 24100292.
- PMID 27046250.
- PMID 24954904.
- PMID 21258367.
- PMID 23332761.
- PMID 18439900.
- PMID 14651849.
- PMID 28743755.
- ^ Jimenez SA, Cronin PM, Koenig AS, et al. (15 February 2012). Varga J, Talavera F, Goldberg E, Mechaber AJ, Diamond HS (eds.). "Scleroderma". Medscape Reference. WebMD. Retrieved 5 March 2014.
- ^ Hajj-ali RA (June 2013). "Systemic Sclerosis". Merck Manual Professional. Merck Sharp & Dohme Corp. Retrieved 5 March 2014.
- ^ "Mammalian target of rapamycin (mTOR) inhibitors in solid tumours". Pharmaceutical Journal. Retrieved 2018-10-18.
- S2CID 27952376.
- PMID 19805415.
- S2CID 205506774.
- PMID 29997249.
- S2CID 45618964.
- PMID 28611316.
- PMID 29618945.
- S2CID 184483900.
- S2CID 147704626.
- ^ "mTOR protein interactors". Human Protein Reference Database. Johns Hopkins University and the Institute of Bioinformatics. Archived from the original on 2015-06-28. Retrieved 2010-12-06.
- PMID 10753870.
- PMID 10910062.
- PMID 14970221.
- PMID 12231510.
- PMID 16541103.
- ^ PMID 12747827.
- ^ PMID 12150926.
- ^ PMID 16798736.
- ^ PMID 15854902.
- ^ S2CID 39048740.
- ^ PMID 9465032.
- PMID 15767663.
- S2CID 27706675.
- PMID 15284440.
- PMID 15796538.
- PMID 7822316.
- PMID 10325225.
- PMID 16837165.
- PMID 16631613.
- ^ PMID 16962653.
- PMID 15459249.
- S2CID 24814691.
- S2CID 21552704.
- PMID 12665511.
- ^ PMID 12604610.
- ^ PMID 16603397.
- PMID 16354680.
- ^ PMID 16183647.
- ^ PMID 17043309.
- PMID 10698949.
- PMID 15878852.
- PMID 15772076.
- S2CID 4427427.
- PMID 11914378.
- PMID 15899889.
- PMID 15905173.
- PMID 10567431.
- PMID 15896331.
- PMID 15809305.
- PMID 14679009.
- PMID 16922504.
- PMID 12807916.
- PMID 10660304.
- PMID 15522880.
- PMID 23394946.
- PMID 11853878.
Further reading
- Saxton RA, Sabatini DM (March 2017). "mTOR Signaling in Growth, Metabolism, and Disease". Cell. 168 (6): 960–976. PMID 28283069.
External links
- mTOR+protein at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- "mTOR Signaling Pathway in Pathway Interaction Database". National Cancer Institute. Archived from the original on 2013-03-18. Retrieved 2015-10-18.
- Overview of all the structural information available in the PDB for UniProt: P42345 (Serine/threonine-protein kinase mTOR) at the PDBe-KB.