Nanoarchaeota
| Nanoarchaeota | |
|---|---|

| |
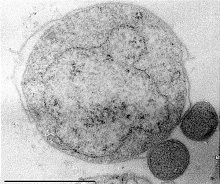
| Nanoarcheotum Nanopusillus acidilobi attached to Acidilobus. | |
| Scientific classification | |
| Domain: | Archaea |
| Superphylum: | DPANN Huber et al. 2002 |
| Phylum: | Nanoarchaeota Huber et al. 2002 |
| Order | |
| |
| Synonyms | |
| |
Nanoarchaeota (Greek, "dwarf or tiny ancient one") is a proposed
Taxonomy
Members of the Nanoarchaeota are associated with different host organisms and environmental conditions.[3] Despite small size, a reduced genome and limited respiration, members of the Nanoarchaeota have unusual metabolic features. For example, N. equitans has a complex and highly developed intercellular communication system.[4]
The phylogeny of the Nanoarchaeota is anchored by its only cultured representative, Nanoarchaeum equitans, which clusters in a separate evolutionary group than other archaea,
The currently accepted taxonomy is based on the List of Prokaryotic names with Standing in Nomenclature (LPSN)[8] and National Center for Biotechnology Information (NCBI).[9]
| Phylogeny of Nanoarchaeota[10][11][12] | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
- Class "Nanoarchaeia" Vazquez-Campos et al. 2021[13] ["Nanoarchaea" Huber et al. 2011;[14] Nanobdellia Kato et al. 2022[15]]
- Order "Jingweiarchaeales" Rao et al. 2023
- Family "Haiyanarchaeaceae" Rao et al. 2023
- Genus ?"Candidatus Haiyanarchaeum" Rao et al. 2023
- "Ca. H. thermophilum" Rao et al. 2023
- Genus ?"Candidatus Haiyanarchaeum" Rao et al. 2023
- Family "Jingweiarchaeaceae" Rao et al. 2023
- Genus ?"Candidatus Jingweiarchaeum" Rao et al. 2023
- "Ca. J. tengchongense" Rao et al. 2023
- Genus ?"Candidatus Jingweiarchaeum" Rao et al. 2023
- Family "Haiyanarchaeaceae" Rao et al. 2023
- Order "Nanoarchaeales" Huber et al. 2011[14] [Nanobdellales Kato et al. 2022[15]]
- Family "Nanoarchaeaceae" Huber et al. 2011[14]
- Genus "Nanoarchaeum" Huber et al. 2002[2]
- "N. equitans" Huber et al. 2002[2]
- Genus "
- Family "Nanopusillaceae" Huber et al. 2011[14] [Nanobdellaceae Kato et al. 2022[15]]
- Family "Nanoarchaeaceae" Huber et al. 2011[14]
- Order "Tiddalikarchaeales" Vazquez-Campos et al. 2021[13]
- Order "Parvarchaeales" Rinke et al. 2020[18]
- Family "Parvarchaeaceae" Rinke et al. 2020[18] ["Acidifodinimicrobiaceae" Luo et al. 2020[19]]
- Order "Jingweiarchaeales" Rao et al. 2023
Characteristics
Cells of N. equitans are spherical with a diameter of approximately 400 nm,[2] and have a very short and compact DNA sequence with the entire genome containing only 490,885 base pairs.[6] While they have the genetic code to carry out processing and repair, they cannot carry out certain biosynthetic and metabolic processes such as lipid, amino-acid, cofactor, or nucleotide synthesis.[6] Due to its limited machinery, it is an obligate parasite, the only one known in the Archaea.[6] Because of their unusual ss rRNA sequences, they are difficult to detect using standard polymerase chain reaction methods.[21] Cells of N. equitans contain a normal S-layer with sixfold symmetry with a 15 nm lattice constant.[21]
Genome structure
Small cells between 100 and 400 nm in diameter and highly streamlined genomes of 0.491-0.606 Mbp characterize nanoarchaeotes.
Habitat
Nanoarchaeotes are obligate symbionts that grow attached to an archaeal host known as
Metabolism
Although much of the metabolism of members of the Nanoarchaeota is unknown, its host is an autotroph that grows on elemental sulphur as an
See also
References
- ^ See the NCBI webpage on Nanoarchaeota. Data extracted from the "NCBI taxonomy resources". National Center for Biotechnology Information. Retrieved 2007-03-19.
- ^ S2CID 4395094.
- ^ PMID 26341207.
- PMID 30223889.
- S2CID 3801477.
- ^ PMID 14566062.
- )
- ^ J.P. Euzéby. "Phylum "Candidatus Nanoarchaeota"". List of Prokaryotic names with Standing in Nomenclature (LPSN). Retrieved 2021-11-17.
- ^ Sayers; et al. "Nanoarchaeota". National Center for Biotechnology Information (NCBI) taxonomy database. Retrieved 2021-06-05.
- ^ "GTDB release 08-RS214". Genome Taxonomy Database. Retrieved 6 December 2021.
- ^ "ar53_r214.sp_label". Genome Taxonomy Database. Retrieved 10 May 2023.
- ^ "Taxon History". Genome Taxonomy Database. Retrieved 6 December 2021.
- ^ PMID 34737728.
- ^ ISBN 978-1-118-96060-8.
- ^ S2CID 251720962.)
{{cite journal}}: CS1 maint: numeric names: authors list (link - ^ S2CID 52178746.
- ^ PMID 27378076.
- ^ S2CID 231984712.
- ^ PMID 33408709.
- ^ PMID 20421484.
- ^ S2CID 4395094.
- ^ "Nanoarchaeota - an overview | ScienceDirect Topics". www.sciencedirect.com. Retrieved 2023-04-08.
- ^ "Nanoarchaeota - an overview | ScienceDirect Topics". www.sciencedirect.com. Retrieved 2023-04-08.
- ^ "Nanoarchaeota - an overview | ScienceDirect Topics". www.sciencedirect.com. Retrieved 2023-04-08.
- ISBN 978-0-387-30743-5, retrieved 2023-04-08
- ^ ISBN 978-3-642-11274-4, retrieved 2023-04-08
Further reading
- Clingenpeel, Scott; Kan, Jinjun; Macur, Richard E.; Woyke, Tanja; et al. (11 September 2013). "Yellowstone Lake Nanoarchaeota". Frontiers in Microbiology. 4: 274. PMID 24062731.
- Hohn, MJ; Hedlund BP; Huber H (2002). "Detection of 16S rDNA sequences representing the novel phylum 'Nanoarchaeota': indication for a wide distribution in high temperature biotopes". Syst. Appl. Microbiol. 25 (4): 551–554. PMID 12583716.
- Huber, H; Hohn MJ; Rachel R; Fuchs T; et al. (2002). "A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont". Nature. 417 (6884): 63–67. S2CID 4395094.
- Stackebrandt, E; Frederiksen W; Garrity GM; Grimont PA; et al. (2002). "Report of the ad hoc committee for the re-evaluation of the species definition in bacteriology". Int. J. Syst. Evol. Microbiol. 52 (Pt 3): 1043–1047. PMID 12054223.
- Christensen, H; Bisgaard M; Frederiksen W; Mutters R; et al. (2001). "Is characterization of a single isolate sufficient for valid publication of a new genus or species? Proposal to modify recommendation 30b of the Bacteriological Code (1990 Revision)". Int. J. Syst. Evol. Microbiol. 51 (Pt 6): 2221–5. PMID 11760965.
- Gurtler, V; Mayall BC (2001). "Genomic approaches to typing, taxonomy and evolution of bacterial isolates". Int. J. Syst. Evol. Microbiol. 51 (Pt 1): 3–16. PMID 11211268.
- Dalevi, D; Hugenholtz P; Blackall LL (2001). "A multiple-outgroup approach to resolving division-level phylogenetic relationships using 16S rDNA data". Int. J. Syst. Evol. Microbiol. 51 (Pt 2): 385–91. PMID 11321083.
- Keswani, J; Whitman WB (2001). "Relationship of 16S rRNA sequence similarity to DNA hybridization in prokaryotes". Int. J. Syst. Evol. Microbiol. 51 (Pt 2): 667–78. PMID 11321113.
- Young, JM (2001). "Implications of alternative classifications and horizontal gene transfer for bacterial taxonomy". Int. J. Syst. Evol. Microbiol. 51 (Pt 3): 945–53. PMID 11411719.
- Christensen, H; Angen O; Mutters R; Olsen JE; et al. (2000). "DNA-DNA hybridization determined in micro-wells using covalent attachment of DNA". Int. J. Syst. Evol. Microbiol. 50 (3): 1095–102. PMID 10843050.
- Xu, HX; Kawamura Y; Li N; Zhao L; et al. (2000). "A rapid method for determining the G+C content of bacterial chromosomes by monitoring fluorescence intensity during DNA denaturation in a capillary tube". Int. J. Syst. Evol. Microbiol. 50 (4): 1463–9. PMID 10939651.
- Young, JM (2000). "Suggestions for avoiding on-going confusion from the Bacteriological Code". Int. J. Syst. Evol. Microbiol. 50 (4): 1687–9. PMID 10939677.
- Hansmann, S; Martin W (2000). "Phylogeny of 33 ribosomal and six other proteins encoded in an ancient gene cluster that is conserved across prokaryotic genomes: influence of excluding poorly alignable sites from analysis". Int. J. Syst. Evol. Microbiol. 50 (4): 1655–63. PMID 10939673.
- Tindall, BJ (1999). "Proposal to change the Rule governing the designation of type strains deposited under culture collection numbers allocated for patent purposes". Int. J. Syst. Bacteriol. 49 (3): 1317–1319. PMID 10490293.
- Tindall, BJ (1999). "Proposal to change Rule 18a, Rule 18f and Rule 30 to limit the retroactive consequences of changes accepted by the ICSB". Int. J. Syst. Bacteriol. 49 (3): 1321–1322. PMID 10425797.
- Tindall, BJ (1999). "Misunderstanding the Bacteriological Code". Int. J. Syst. Bacteriol. 49 (3): 1313–1316. PMID 10425796.
- Tindall, BJ (1999). "Proposals to update and make changes to the Bacteriological Code". Int. J. Syst. Bacteriol. 49 (3): 1309–1312. PMID 10425795.
- Palys, T; Nakamura LK; Cohan FM (1997). "Discovery and classification of ecological diversity in the bacterial world: the role of DNA sequence data". Int. J. Syst. Bacteriol. 47 (4): 1145–1156. PMID 9336922.
- Euzeby, JP (1997). "List of Bacterial Names with Standing in Nomenclature: a folder available on the Internet". Int. J. Syst. Bacteriol. 47 (2): 590–592. PMID 9103655.
- Clayton, RA; Sutton G; Hinkle PS Jr; Bult C; Fields C (1995). "Intraspecific variation in small-subunit rRNA sequences in GenBank: why single sequences may not adequately represent prokaryotic taxa". Int. J. Syst. Bacteriol. 45 (3): 595–599. PMID 8590690.
- Murray, RG; Schleifer KH (1994). "Taxonomic notes: a proposal for recording the properties of putative taxa of procaryotes". Int. J. Syst. Bacteriol. 44 (1): 174–176. PMID 8123559.
- Winker, S; Woese CR (1991). "A definition of the domains Archaea, Bacteria and Eucarya in terms of small subunit ribosomal RNA characteristics". Syst. Appl. Microbiol. 14 (4): 305–10. PMID 11540071.
- Woese, CR; Kandler O; Wheelis ML (1990). "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proc. Natl. Acad. Sci. USA. 87 (12): 4576–4579. PMID 2112744.
- Achenbach-Richter, L; Woese CR (1988). "The ribosomal gene spacer region in archaebacteria". Syst. Appl. Microbiol. 10 (3): 211–4. PMID 11542149.
- McGill, TJ; Jurka J; Sobieski JM; Pickett MH; et al. (1986). "Characteristic archaebacterial 16S rRNA oligonucleotides". Syst. Appl. Microbiol. 7 (2–3): 194–7. PMID 11542064.
- Woese, CR; Olsen GJ; Hahn, C. M.; Zillig, W; et al. (1984). "The phylogenetic relationships of three sulfur dependent archaebacteria". Syst. Appl. Microbiol. 5: 97–105. PMID 11541975.
- Woese, CR; Fox GE (1977). "Phylogenetic structure of the prokaryotic domain: the primary kingdoms". Proc. Natl. Acad. Sci. USA. 74 (11): 5088–5090. PMID 270744.
