Gut microbiota

Gut microbiota, gut microbiome, or gut flora are the
The microbial composition of the gut microbiota varies across regions of the digestive tract. The
Overview

In humans, the gut microbiota has the largest numbers and species of bacteria compared to other areas of the body.[9] The approximate number of bacteria composing the gut microbiota is about 1013–1014.[10] In humans, the gut flora is established at one to two years after birth, by which time the intestinal epithelium and the intestinal mucosal barrier that it secretes have co-developed in a way that is tolerant to, and even supportive of, the gut flora and that also provides a barrier to pathogenic organisms.[11][12]
The relationship between some gut microbiota and humans is not merely
The composition of human gut microbiota changes over time, when the diet changes, and as overall health changes.
Classifications
The microbial composition of the gut microbiota varies across the digestive tract. In the
Over 99% of the bacteria in the gut are
Many species in the gut have not been studied outside of their hosts because they cannot be cultured.[16][7][21] While there are a small number of core microbial species shared by most individuals, populations of microbes can vary widely.[22] Within an individual, their microbial populations stay fairly constant over time, with some alterations occurring due to changes in lifestyle, diet and age.[6][23] The Human Microbiome Project has set out to better describe the microbiota of the human gut and other body locations.[citation needed]
The four dominant bacterial phyla in the human gut are Bacillota (Firmicutes), Bacteroidota, Actinomycetota, and Pseudomonadota.[24] Most bacteria belong to the genera Bacteroides, Clostridium, Faecalibacterium,[6][7] Eubacterium, Ruminococcus, Peptococcus, Peptostreptococcus, and Bifidobacterium.[6][7] Other genera, such as Escherichia and Lactobacillus, are present to a lesser extent.[6] Species from the genus Bacteroides alone constitute about 30% of all bacteria in the gut, suggesting that this genus is especially important in the functioning of the host.[16]
Fungal genera that have been detected in the gut include
Archaea constitute another large class of gut flora which are important in the metabolism of the bacterial products of fermentation.
Industrialization is associated with changes in the microbiota and the reduction of diversity could drive certain species to extinction; in 2018, researchers proposed a biobank repository of human microbiota.[27]
Enterotype
An enterotype is a classification of living organisms based on its bacteriological ecosystem in the human gut microbiome not dictated by age, gender, body weight, or national divisions.[28] There are indications that long-term diet influences enterotype.[29] Three human enterotypes have been proposed,[28][30] but their value has been questioned.[31]
Composition
Bacteriome
Stomach
Due to the high acidity of the
Intestines
| Bacteria commonly found in the human colon[34] | |
| Bacterium | Incidence (%) |
|---|---|
| Bacteroides fragilis | 100 |
Bacteroides melaninogenicus
|
100 |
| Bacteroides oralis | 100 |
| Enterococcus faecalis | 100 |
| Escherichia coli | 100 |
| Enterobacter sp. | 40–80 |
| Klebsiella sp. | 40–80 |
| Bifidobacterium bifidum | 30–70 |
| Staphylococcus aureus | 30–50 |
| Lactobacillus | 20–60 |
| Clostridium perfringens | 25–35 |
| Proteus mirabilis | 5–55 |
| Clostridium tetani | 1–35 |
| Clostridium septicum | 5–25 |
| Pseudomonas aeruginosa | 3–11 |
| Salmonella enterica | 3–7 |
Faecalibacterium prausnitzii
|
?common |
| Peptostreptococcus sp. | ?common |
| Peptococcus sp. | ?common |
The small intestine contains a trace amount of microorganisms due to the proximity and influence of the stomach. Gram-positive cocci and rod-shaped bacteria are the predominant microorganisms found in the small intestine.[5] However, in the distal portion of the small intestine alkaline conditions support gram-negative bacteria of the Enterobacteriaceae.[5] The bacterial flora of the small intestine aid in a wide range of intestinal functions. The bacterial flora provide regulatory signals that enable the development and utility of the gut. Overgrowth of bacteria in the small intestine can lead to intestinal failure.[35] In addition the large intestine contains the largest bacterial ecosystem in the human body.[5] About 99% of the large intestine and feces flora are made up of obligate anaerobes such as Bacteroides and Bifidobacterium.[36] Factors that disrupt the microorganism population of the large intestine include antibiotics, stress, and parasites.[5]
Bacteria make up most of the flora in the
This fact makes feces an ideal source of gut flora for any tests and experiments by extracting the nucleic acid from fecal specimens, and bacterial 16S rRNA gene sequences are generated with bacterial primers. This form of testing is also often preferable to more invasive techniques, such as biopsies.Five
Research suggests that the relationship between gut
Mycobiome
Fungi and protists also make up a part of the gut flora, but less is known about their activities.[43]
Virome
The human virome is mostly bacteriophages.[44]
Variation
Age
There are common patterns of microbiome composition evolution during life.[45] In general, the diversity of microbiota composition of fecal samples is significantly higher in adults than in children, although interpersonal differences are higher in children than in adults.[46] Much of the maturation of microbiota into an adult-like configuration happens during the first three years of life.[46]
As the microbiome composition changes, so does the composition of bacterial proteins produced in the gut. In adult microbiomes, a high prevalence of enzymes involved in fermentation, methanogenesis and the metabolism of arginine, glutamate, aspartate and lysine have been found. In contrast, in infant microbiomes the dominant enzymes are involved in cysteine metabolism and fermentation pathways.[46]
Geography
Gut microbiome composition depends on the geographic origin of populations. Variations in a trade-off of Prevotella, the representation of the urease gene, and the representation of genes encoding glutamate synthase/degradation or other enzymes involved in amino acids degradation or vitamin biosynthesis show significant differences between populations from the US, Malawi, or Amerindian origin.[46]
The US population has a high representation of enzymes encoding the degradation of
Further studies have indicated a large difference in the composition of microbiota between European and rural African children. The fecal bacteria of children from Florence were compared to that of children from the small rural village of Boulpon in Burkina Faso. The diet of a typical child living in this village is largely lacking in fats and animal proteins and rich in polysaccharides and plant proteins. The fecal bacteria of European children were dominated by Firmicutes and showed a marked reduction in biodiversity, while the fecal bacteria of the Boulpon children was dominated by Bacteroidetes. The increased biodiversity and different composition of the gut microbiome in African populations may aid in the digestion of normally indigestible plant polysaccharides and also may result in a reduced incidence of non-infectious colonic diseases.[47]
On a smaller scale, it has been shown that sharing numerous common environmental exposures in a family is a strong determinant of individual microbiome composition. This effect has no genetic influence and it is consistently observed in culturally different populations.[46]
Malnourishment
Malnourished children have less mature and less diverse gut microbiota than healthy children, and changes in the microbiome associated with nutrient scarcity can in turn be a pathophysiological cause of malnutrition.[48][49] Malnourished children also typically have more potentially pathogenic gut flora, and more yeast in their mouths and throats.[50] Altering diet may lead to changes in gut microbiota composition and diversity.[51]
Race and ethnicity
Researchers with the American Gut Project and Human Microbiome Project found that twelve microbe families varied in abundance based on the race or ethnicity of the individual. The strength of these associations is limited by the small sample size: the American Gut Project collected data from 1,375 individuals, 90% of whom were white.[52] The Healthy Life in an Urban Setting (HELIUS) study in Amsterdam found that those of Dutch ancestry had the highest level of gut microbiota diversity, while those of South Asian and Surinamese descent had the lowest diversity. The study results suggested that individuals of the same race or ethnicity have more similar microbiomes than individuals of different racial backgrounds.[52]
Socioeconomic status
As of 2020, at least two studies have demonstrated a link between an individual's socioeconomic status (SES) and their gut microbiota. A study in Chicago found that individuals in higher SES neighborhoods had greater microbiota diversity. People from higher SES neighborhoods also had more abundant Bacteroides bacteria. Similarly, a study of twins in the United Kingdom found that higher SES was also linked with a greater gut diversity.[52]
Acquisition in human infants
The establishment of a gut flora is crucial to the health of an adult, as well as the functioning of the gastrointestinal tract.[53] In humans, a gut flora similar to an adult's is formed within one to two years of birth as microbiota are acquired through parent-to-child transmission and transfer from food, water, and other environmental sources.[54][11]

The traditional view of the gastrointestinal tract of a normal fetus is that it is sterile, although this view has been challenged in the past few years.[timeframe?][55] Multiple lines of evidence have begun to emerge that suggest there may be bacteria in the intrauterine environment. In humans, research has shown that microbial colonization may occur in the fetus[56] with one study showing Lactobacillus and Bifidobacterium species were present in placental biopsies.[57] Several rodent studies have demonstrated the presence of bacteria in the amniotic fluid and placenta, as well as in the meconium of babies born by sterile cesarean section.[58][59] In another study, researchers administered a culture of bacteria orally to pregnant mice, and detected the bacteria in the offspring, likely resulting from transmission between the digestive tract and amniotic fluid via the blood stream.[60] However, researchers caution that the source of these intrauterine bacteria, whether they are alive, and their role, is not yet understood.[61][57]
During birth and rapidly thereafter, bacteria from the mother and the surrounding environment colonize the infant's gut.[11] The exact sources of bacteria are not fully understood, but may include the birth canal, other people (parents, siblings, hospital workers), breastmilk, food, and the general environment with which the infant interacts.[62] Research has shown that the microbiome of babies born vaginally differs significantly from that of babies delivered by caesarean section and that vaginally born babies got most of their gut bacteria from their mother, while the microbiota of babies born by caesarean section had more bacteria associated with hospital environments.[63]
During the first year of life, the composition of the gut flora is generally simple and changes a great deal with time and is not the same across individuals.
Caesarean section,
Functions
When the study of gut flora began in 1995,[68] it was thought to have three key roles: direct defense against pathogens, fortification of host defense by its role in developing and maintaining the intestinal epithelium and inducing antibody production there, and metabolizing otherwise indigestible compounds in food. Subsequent work discovered its role in training the developing immune system, and yet further work focused on its role in the gut–brain axis.[69]
Direct inhibition of pathogens
The gut flora community plays a direct role in defending against pathogens by fully colonising the space, making use of all available nutrients, and by secreting compounds known as
Development of enteric protection and immune system
In humans, a gut flora similar to an adult's is formed within one to two years of birth.
The human
The immune system can also be altered due to the gut bacteria's ability to produce
Metabolism
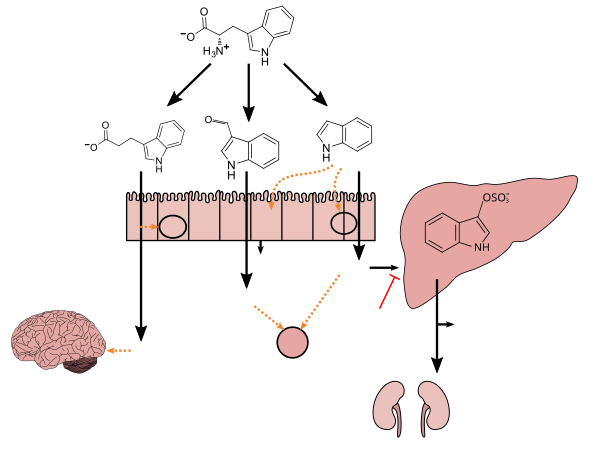
Tryptophan metabolism by
human gastrointestinal microbiota ( )activated charcoal), an intestinal sorbent that is taken by mouth, adsorbs indole, in turn decreasing the concentration of indoxyl sulfate in blood plasma.[76] |
Without gut flora, the human body would be unable to utilize some of the undigested
Bacteria turn carbohydrates they ferment into short-chain fatty acids by a form of fermentation called saccharolytic fermentation.[40] Products include acetic acid, propionic acid and butyric acid.[7][40] These materials can be used by host cells, providing a major source of energy and nutrients.[40] Gases (which are involved in signaling[80] and may cause flatulence) and organic acids, such as lactic acid, are also produced by fermentation.[7] Acetic acid is used by muscle, propionic acid facilitates liver production of ATP, and butyric acid provides energy to gut cells.[40]
Gut flora also synthesize vitamins like
Gut microbiota also serve as a source of vitamins K and B12, which are not produced by the body or produced in little amount.[82][83]
Cellulose degradation
Bacteria that degrade cellulose (such as Ruminococcus) are prevalent among great apes, ancient human societies, hunter-gatherer communities, and even modern rural populations. However, they are rare in industrialized societies. Human-associated strains have acquired genes that can degrade specific plant fibers such as maize, rice, and wheat. Bacterial strains found in primates can also degrade chitin, a polymer abundant in insects, which are part of the diet of many nonhuman primates. The decline of these bacteria in the human gut were likely influenced by the shift toward western lifestyles.[84]
Pharmacomicrobiomics
The human
Apart from carbohydrates, gut microbiota can also metabolize other xenobiotics such as drugs, phytochemicals, and food toxicants. More than 30 drugs have been shown to be metabolized by gut microbiota.[89] The microbial metabolism of drugs can sometimes inactivate the drug.[90]
Contribution to drug metabolism
The gut microbiota is an enriched community that contains diverse genes with huge biochemical capabilities to modify drugs, especially those taken by mouth.[91] Gut microbiota can affect drug metabolism via direct and indirect mechanisms.[92] The direct mechanism is mediated by the microbial enzymes that can modify the chemical structure of the administered drugs.[93] Conversely, the indirect pathway is mediated by the microbial metabolites which affect the expression of host metabolizing enzymes such as cytochrome P450.[94][92] The effects of the gut microbiota on the pharmacokinetics and bioavailability of the drug have been investigated a few decades ago.[95][96][97] These effects can be varied; it could activate the inactive drugs such as lovastatin,[98] inactivate the active drug such as digoxin[99] or induce drug toxicity as in irinotecan.[100] Since then, the impacts of the gut microbiota on the pharmacokinetics of many drugs were heavily studied.[101][91]
The human gut microbiota plays a crucial role in modulating the effect of the administered drugs on the human. Directly, gut microbiota can synthesize and release a series of enzymes with the capability to metabolize drugs such as microbial biotransformation of L-dopa by decarboxylase and dehydroxylase enzymes.[93] On the contrary, gut microbiota may also alter the metabolism of the drugs by modulating the host drug metabolism. This mechanism can be mediated by microbial metabolites or by modifying host metabolites which in turn change the expression of host metabolizing enzymes.[94]
A large number of studies have demonstrated the metabolism of over 50 drugs by the gut microbiota.[101][92] For example, lovastatin (a cholesterol-lowering agent) which is a lactone prodrug is partially activated by the human gut microbiota forming active acid hydroxylated metabolites.[98] Conversely, digoxin (a drug used to treat Congestive Heart Failure) is inactivated by a member of the gut microbiota (i.e. Eggerthella lanta).[102] Eggerthella lanta has a cytochrome-encoding operon up-regulated by digoxin and associated with digoxin-inactivation.[102] Gut microbiota can also modulate the efficacy and toxicity of chemotherapeutic agents such as irinotecan.[103] This effect is derived from the microbiome-encoded β-glucuronidase enzymes which recover the active form of the irinotecan causing gastrointestinal toxicity.[104]
Secondary metabolites
This microbial community in the gut has a huge biochemical capability to produce distinct secondary metabolites that are sometimes produced from the metabolic conversion of dietary foods such as fibers, endogenous biological compounds such as indole or bile acids.[105][106][107] Microbial metabolites especially short chain fatty acids (SCFAs) and secondary bile acids (BAs) play important roles for the human in health and disease states.[108][109][110]
One of the most important bacterial metabolites produced by the gut microbiota is secondary bile acids (BAs).[107] These metabolites are produced by the bacterial biotransformation of the primary bile acids such as cholic acid (CA) and chenodeoxycholic acid (CDCA) into secondary bile acids (BAs) lithocholic acid (LCA) and deoxy cholic acid (DCA) respectively.[111] Primary bile acids which are synthesized by hepatocytes and stored in the gall bladder possess hydrophobic characters. These metabolites are subsequently metabolized by the gut microbiota into secondary metabolites with increased hydrophobicity.[111] Bile salt hydrolases (BSH) which are conserved across gut microbiota phyla such as Bacteroides, Firmicutes, and Actinobacteria responsible for the first step of secondary bile acids metabolism.[111] Secondary bile acids (BAs) such as DCA and LCA have been demonstrated to inhibit both Clostridium difficile germination and outgrowth.[110]
Dysbiosis
The gut microbiota is important for maintaining homeostasis in the intestine. Development of
Gut–brain axis
The gut–brain axis is the biochemical signaling that takes place between the
A
Alterations in microbiota balance
Effects of antibiotic use
Altering the numbers of gut bacteria, for example by taking
Changing the numbers and species of gut microbiota can reduce the body's ability to ferment carbohydrates and metabolize bile acids and may cause diarrhea. Carbohydrates that are not broken down may absorb too much water and cause runny stools, or lack of SCFAs produced by gut microbiota could cause diarrhea.[7]
A reduction in levels of native bacterial species also disrupts their ability to inhibit the growth of harmful species such as C. difficile and
The composition of the gut microbiome also changes in severe illnesses, due not only to antibiotic use but also to such factors as
Antibiotics alter the population of the microbiota in the gastrointestinal tract, and this may change the intra-community metabolic interactions, modify caloric intake by using carbohydrates, and globally affects host metabolic, hormonal and immune homeostasis.[120]
There is reasonable evidence that taking probiotics containing Lactobacillus species may help prevent antibiotic-associated diarrhea and that taking probiotics with Saccharomyces (e.g., Saccharomyces boulardii ) may help to prevent Clostridium difficile infection following systemic antibiotic treatment.[121]
Pregnancy
The gut microbiota of a woman changes as
Probiotics, prebiotics, synbiotics, and pharmabiotics
The term "pharmabiotics" is used in various ways, to mean:
There is some evidence that treatment with some probiotic strains of bacteria may be effective in
- Bifidobacterium breve
- Bifidobacterium infantis
- Enterococcus faecium
- Lactobacillus plantarum
- Lactobacillus reuteri
- Lactobacillus rhamnosus
- Lactobacillus salivarius
- Propionibacterium freudenreichii
- Saccharomyces boulardii
- Escherichia coli Nissle 1917
- Streptococcus thermophilus[132][133][134]
Fecal floatation
Feces of about 10–15% of people consistently floats in toilet water ('floaters'), while the rest produce feces that sinks ('sinkers') and production of gas causes feces to float.[135] While conventional mice often produce 'floaters', gnotobiotic germfree mice no gut microbiota (bred in germfree isolator) produce 'sinkers', and gut microbiota colonization in germfree mice leads to food transformation to microbial biomass and enrichment of multiple gasogenic bacterial species that turns the 'sinkers' into 'floaters'.[136]
Research
Tests for whether non-antibiotic drugs may impact human gut-associated bacteria were performed by in vitro analysis on more than 1000 marketed drugs against 40 gut bacterial strains, demonstrating that 24% of the drugs inhibited the growth of at least one of the bacterial strains.[137]
Effects of exercise
Gut microbiota and exercise have recently been shown to be interconnected. Both moderate and intense exercise are typically part of the training regimen of endurance athletes, but they exert different effects on health. The interconnection between gut microbiota and endurance sports depends upon exercise intensity and training status.[138]
Role in disease
Bacteria in the digestive tract can contribute to and be affected by disease in various ways. The presence or overabundance of some kinds of bacteria may contribute to inflammatory disorders such as
Ulcers
Bowel perforation
Normally-
Inflammatory bowel diseases
The two main types of inflammatory bowel diseases, Crohn's disease and ulcerative colitis, are chronic inflammatory disorders of the gut; the causes of these diseases are unknown and issues with the gut flora and its relationship with the host have been implicated in these conditions.[14][141][142][143] Additionally, it appears that interactions of gut flora with the gut–brain axis have a role in IBD, with physiological stress mediated through the hypothalamic–pituitary–adrenal axis driving changes to intestinal epithelium and the gut flora in turn releasing factors and metabolites that trigger signaling in the enteric nervous system and the vagus nerve.[4]
The diversity of gut flora appears to be significantly diminished in people with inflammatory bowel diseases compared to healthy people; additionally, in people with ulcerative colitis, Proteobacteria and Actinobacteria appear to dominate; in people with Crohn's, Enterococcus faecium and several Proteobacteria appear to be over-represented.[4]
There is reasonable evidence that correcting gut flora imbalances by taking probiotics with
Irritable bowel syndrome
Asthma
With asthma, two hypotheses have been posed to explain its rising prevalence in the developed world. The
Diabetes mellitus type 1
The connection between the gut microbiota and
Obesity and metabolic syndrome
The gut flora have been implicated in obesity and
Additionally, the liver plays a dominant role in
Just as gut flora can function in a feedback loop that can drive the development of obesity, there is evidence that restricting intake of calories (i.e., dieting) can drive changes to the composition of the gut flora.[143]
Other animals
The composition of the human gut microbiome is similar to that of the other great apes. However, humans' gut biota has decreased in diversity and changed in composition since our evolutionary split from Pan.[153] Humans display increases in Bacteroidetes, a bacterial phylum associated with diets high in animal protein and fat, and decreases in Methanobrevibacter and Fibrobacter, groups that ferment complex plant polysaccharides.[153] These changes are the result of the combined dietary, genetic, and cultural changes humans have undergone since evolutionary divergence from Pan.[citation needed]
In addition to humans and vertebrates, some insects also have complex and diverse gut microbiota that play key nutritional roles.
For more than 51 years it has been known that the administration of low doses of antibacterial agents promotes the growth of farm animals to increase weight gain.[120]
In a study carried out on
See also
- Colonisation resistance
- List of human flora
- List of microbiota species of the lower reproductive tract of women
- Skin flora
- Verotoxin-producing Escherichia coli
Notes
References
- S2CID 216108564.
- ^ PMID 23692388.
- PMID 23670539.
- ^ ISBN 978-0-12-799922-7.
- ^ OCLC 886600661.
- ^ S2CID 38767655.
- ^ PMID 15123074.
- ^ PMID 7359576.
- ^ PMID 24729765.
- PMID 17943116.
- ^ S2CID 22798964.
- ^ S2CID 25878594.
- ^ PMID 24892638.
- ^ PMID 27195115.
- ^ PMID 27413138.
- ^ PMID 16701579.
- PMID 27039196.
- PMID 37524974.
- PMID 22972295.
- PMID 20203603.
- PMID 12473298.
- PMID 19601958.
- ^ PMID 16819463.
- PMID 24388028.
- ^ PMID 23899327.
- S2CID 3098136.
- S2CID 52919917.
- ^ PMID 21508958.
- PMID 21885731.
- ^ Zimmer, Carl (April 20, 2011). "Bacteria Divide People Into 3 Types, Scientists Say". The New York Times. Retrieved April 21, 2011.
a group of scientists now report just three distinct ecosystems in the guts of people they have studied.
- PMID 25299329.
- ^ PMID 26137299.
- PMID 30831083.
- ^ Todar, Kenneth (2012). "The Normal Bacterial Flora of Humans". Todar's Online Textbook of Bacteriology. Retrieved June 25, 2016.
- S2CID 16904501.
- S2CID 241261974.
- ^ a b "The normal gut flora" (PDF) (slideshow). 2004. Archived from the original (PDF) on 2004-05-26. Retrieved 2023-01-02 – via University of Glasgow.
- PMID 26963713.
- ^ PMID 15894105.
- ^ .
- PMID 23831042.
- S2CID 23329156.
- PMID 29178920.
- PMID 26257129.
- PMID 21617937.
- ^ PMID 22699611.
- PMID 20679230.
- PMID 27086883.
- S2CID 20381329.
- PMID 25153531.
- ^ PMID 25103109.
- ^ PMID 31635533.
- PMID 22606315.
- PMID 29282061.
- PMID 28454555.
- PMID 23332725.
- ^ PMID 25578246.
- S2CID 43438656.
- PMID 27001291.
- PMID 18281199.
- PMID 28454555.
- S2CID 205859933.
- ^ "Babies' gut bacteria affected by delivery method: Vaginal delivery promotes mother's gut bacteria in babies' gut". ScienceDaily. 18 September 2019. Archived from the original on 24 November 2021. Retrieved 31 May 2022.
- PMID 17259094.
- S2CID 258302702.
- S2CID 10316311.
- PMID 27306663.
- PMID 7782892.
- ^ PMID 24370461.
- ^ S2CID 54622675.
- PMID 26947707.
- PMID 21975936.
- ^ S2CID 3351351.
- ^ S2CID 4461492.
- ^ PMID 27474437.
- ^ PMID 27102537.
Lactobacillus spp. convert tryptophan to indole-3-aldehyde (I3A) through unidentified enzymes [125]. Clostridium sporogenes convert tryptophan to IPA [6], likely via a tryptophan deaminase. ... IPA also potently scavenges hydroxyl radicals
Table 2: Microbial metabolites: their synthesis, mechanisms of action, and effects on health and disease
Figure 1: Molecular mechanisms of action of indole and its metabolites on host physiology and disease - PMID 19234110.
Production of IPA was shown to be completely dependent on the presence of gut microflora and could be established by colonization with the bacterium Clostridium sporogenes.
IPA metabolism diagram - ^ "3-Indolepropionic acid". Human Metabolome Database. University of Alberta. Retrieved 12 June 2018.
- S2CID 6630247.
[Indole-3-propionic acid (IPA)] has previously been identified in the plasma and cerebrospinal fluid of humans, but its functions are not known. ... In kinetic competition experiments using free radical-trapping agents, the capacity of IPA to scavenge hydroxyl radicals exceeded that of melatonin, an indoleamine considered to be the most potent naturally occurring scavenger of free radicals. In contrast with other antioxidants, IPA was not converted to reactive intermediates with pro-oxidant activity.
- S2CID 224824871.
- PMID 24861948.
- S2CID 8740364.
- ^ "The Microbiome". Tufts Now. 2013-09-17. Retrieved 2020-12-09.
- PMC 7615765.
- ^ PMID 24785449.
- ^ PMID 22411464.
The composition of the microbiome varies by anatomical site (Figure 1). The primary determinant of community composition is anatomical location: interpersonal variation is substantial23,24 and is higher than the temporal variability seen at most sites in a single individual25.
- PMID 25774294.
- S2CID 73413138.
- PMID 18682282.
- PMID 23869020.
- ^ PMID 28642381.
- ^ PMID 26972811.
- ^ PMID 31196984.
- ^ PMID 33312848.
- PMID 1204496.
- PMID 6836275.
- PMID 6804597.
- ^ S2CID 7524335.
- PMID 23869020.
- PMID 34074730.
- ^ PMID 18682282.
- ^ PMID 29761785.
- S2CID 9654170.
- S2CID 26655779.
- S2CID 8562345.
- S2CID 13979774.
- ^ S2CID 252970168.
- PMID 32901017.
- S2CID 254132108.
- ^ PMID 28279860.
- ^ PMID 18757757.
- ^ PMID 28187526.
- ^ PMID 36408538.
- PMID 25392516.
- ^ S2CID 21424690.
- ^ PMID 15546686.
- PMID 32642633.
- S2CID 2508836.
- S2CID 40683723.
- ^ PMID 22914093.
- ^ PMID 26845162.
- S2CID 87078157.
- PMID 24912386.
- PMID 21861940.
- PMID 26431716.
- PMID 26604335.
- PMID 27050865.
- PMID 19753318.
- PMID 25196939.
- ^ "Non-prescription therapeutics for IBS: Where are we?". 30 January 2024.
- S2CID 33659736.
- S2CID 205100508.
- PMID 25525379.
- S2CID 26982085.
- PMID 5015442.
- PMID 36302811.
- PMID 29555994.
- PMID 34179053.
- ^ PMID 28209367.
- ^ a b c "Peptic ulcer disease" (PDF). The Johns Hopkins University School of Medicine. 2013. Retrieved 21 October 2020.
- PMID 23395397.
- PMID 27179626.
- ^ PMID 27098727.
- ^ S2CID 206687974.
- ^ PMID 28077947.
- S2CID 27328940.
- S2CID 30078908.
- PMID16648838.
- ^ S2CID 41553220.
- PMID 19366864.
- PMID 26916014.
- PMID 23686721.
- ^ PMID 25368157.
- S2CID 5220210.
- ^ PMID 24487532.
- ^ S2CID 206182668.
- PMID 26655763.
Further reading
- Review articles
- De Preter, Vicky; Hamer, Henrike M; Windey, Karen; Verbeke, Kristin (2011). "The impact of pre- and/or probiotics on human colonic metabolism: Does it affect human health?". Molecular Nutrition & Food Research. 55 (1): 46–57. PMID 21207512.
- Maranduba, Carlos Magno da Costa; De Castro, Sandra Bertelli Ribeiro; Souza, Gustavo Torres de; Rossato, Cristiano; Da Guia, Francisco Carlos; Valente, Maria Anete Santana; Rettore, João Vitor Paes; Maranduba, Claudinéia Pereira; Souza, Camila Maurmann de; Carmo, Antônio Márcio Resende do; MacEdo, Gilson Costa; Silva, Fernando de Sá (2015). "Intestinal Microbiota as Modulators of the Immune System and Neuroimmune System: Impact on the Host Health and Homeostasis". Journal of Immunology Research. 2015: 931574. PMID 25759850.
- Prakash, Satya; Rodes, Laetitia; Coussa-Charley, Michael; Tomaro-Duchesneau, Catherine; Tomaro-Duchesneau, Catherine; Coussa-Charley; Rodes (2011). "Gut microbiota: Next frontier in understanding human health and development of biotherapeutics". Biologics: Targets and Therapy. 5: 71–86. PMID 21847343.
- Wu, G. D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.-Y.; Keilbaugh, S. A.; Bewtra, M.; Knights, D.; Walters, W. A.; Knight, R.; Sinha, R.; Gilroy, E.; Gupta, K.; Baldassano, R.; Nessel, L.; Li, H.; Bushman, F. D.; Lewis, J.D. (2011). "Linking Long-Term Dietary Patterns with Gut Microbial Enterotypes". Science. 334 (6052): 105–108. PMID 21885731.