Ras GTPase
| CDD | cd04138 | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
Ras, from "Rat sarcoma virus", is a family of related proteins that are expressed in all animal cell lineages and organs. All Ras protein family members belong to a class of protein called small GTPase, and are involved in transmitting signals within cells (cellular signal transduction). Ras is the prototypical member of the Ras superfamily of proteins, which are all related in three-dimensional structure and regulate diverse cell behaviours.
When Ras is 'switched on' by incoming signals, it subsequently switches on other proteins, which ultimately turn on genes involved in
Because these signals result in cell growth and division, overactive Ras signaling can ultimately lead to cancer.[1] The three Ras genes in humans (HRAS, KRAS, and NRAS) are the most common oncogenes in human cancer; mutations that permanently activate Ras are found in 20 to 25% of all human tumors and up to 90% in certain types of cancer (e.g., pancreatic cancer).[2] For this reason, Ras inhibitors are being studied as a treatment for cancer and other diseases with Ras overexpression.
History
The first two Ras genes,
The three human ras genes encode extremely similar proteins made up of chains of 188 to 189 amino acids. Their gene symbols are HRAS, NRAS and KRAS, the latter of which produces the K-Ras4A and K-Ras4B isoforms from alternative splicing.[citation needed]
Structure

Ras contains six
The G domain contains five G motifs that bind GDP/GTP directly. The G1 motif, or the P-loop, binds the beta phosphate of GDP and GTP. The G2 motif, also called Switch I or SW1, contains threonine35, which binds the terminal phosphate (γ-phosphate) of GTP and the divalent magnesium ion bound in the active site. The G3 motif, also called Switch II or SW2, has a DXXGQ motif. The D is aspartate57, which is specific for guanine versus adenine binding, and Q is glutamine61, the crucial residue that activates a catalytic water molecule for hydrolysis of GTP to GDP. The G4 motif contains a LVGNKxDL motif, and provides specific interaction to guanine. The G5 motif contains a SAK consensus sequence. The A is alanine146, which provides specificity for guanine rather than adenine.
The two switch motifs, G2 (SW1) and G3 (SW2), are the main parts of the protein that move when GTP is hydrolyzed into GDP. This conformational change by the two switch motifs is what mediates the basic functionality as a molecular switch protein. This GTP-bound state of Ras is the "on" state, and the GDP-bound state is the "off" state. The two switch motifs have a number of conformations when binding GTP or GDP or no nucleotide (when bound to SOS1, which releases the nucleotide).[15]
Ras also binds a magnesium ion which helps to coordinate nucleotide binding.
Function
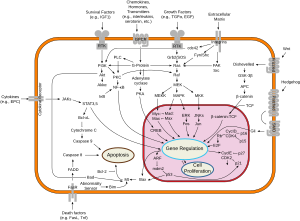
Ras proteins function as binary molecular switches that control intracellular signaling networks. Ras-regulated
Ras activates several pathways, of which the
Activation and deactivation
Ras is a
The process of exchanging the bound nucleotide is facilitated by
GEFs catalyze a "push and pull" reaction which releases GDP from Ras. They insert close to the P-loop and magnesium
The balance between GEF and GAP activity determines the guanine nucleotide status of Ras, thereby regulating Ras activity.
In the GTP-bound conformation, Ras has a high affinity for numerous
Other proteins exist that may change the activity of Ras family proteins. One example is GDI (GDP Disassociation Inhibitor). These function by slowing the exchange of GDP for GTP, thus prolonging the inactive state of Ras family members. Other proteins that augment this cycle may exist.
Membrane attachment
Ras is attached to the
Members
The clinically most notable members of the Ras subfamily are HRAS, KRAS and NRAS, mainly for being implicated in many types of cancer.[18]
However, there are many other members of this subfamily as well:[19]
Ras in cancer
Mutations in the Ras family of
Inappropriate activation
Inappropriate activation of the gene has been shown to play a key role in improper signal transduction, proliferation and malignant transformation.[16]
Mutations in a number of different genes as well as RAS itself can have this effect. Oncogenes such as p210BCR-ABL or the growth receptor erbB are upstream of Ras, so if they are constitutively activated their signals will transduce through Ras.[citation needed]
The
Finally, Ras oncogenes can be activated by point mutations so that the GTPase reaction can no longer be stimulated by GAP – this increases the half life of active Ras-GTP mutants.[23]
Constitutively active Ras
Constitutively active Ras (RasD) is one which contains mutations that prevent GTP hydrolysis, thus locking Ras in a permanently 'On' state.
The most common mutations are found at residue G12 in the
- The glycine to valine mutation at residue 12 renders the GTPase domain of Ras insensitive to inactivation by GAP and thus stuck in the "on state". Ras requires a GAP for inactivation as it is a relatively poor catalyst on its own, as opposed to other G-domain-containing proteins such as the alpha subunit of heterotrimeric G proteins.
- Residue 61[24] is responsible for stabilizing the transition state for GTP hydrolysis. Because enzyme catalysis in general is achieved by lowering the energy barrier between substrate and product, mutation of Q61 to K (Glutamine to Lysine) necessarily reduces the rate of intrinsic Ras GTP hydrolysis to physiologically meaningless levels.
See also "dominant negative" mutants such as S17N and D119N.
Ras-targeted cancer treatments
Another tumor-lysing virus that specifically targets tumor cells with an activated Ras pathway is a type II
In glioblastoma mouse models SHP2 levels were heightened in cancerous brain cells. Inhibiting SHP2 in turn inhibited Ras dephosphorylation. This reduced tumor sizes and accompanying rise in survival rates.[29][30]
Other strategies have attempted to manipulate the regulation of the above-mentioned localization of Ras.
A novel inhibitor finding strategy for mutated Ras molecules was described in.[32] The Ras mutations in the 12th residue position inhibit the bound of the regulatory GAP molecule to the mutated Ras, causing uncontrolled cell growth. The novel strategy proposes finding small glue molecules, which attach the mutated Ras to the GAP, prohibiting uncontrolled cell growth and restoring the normal function. For this goal a theoretical Ras-GAP conformation was designed with a several Å gap between the molecules, and a high-throughput in silico docking was performed for finding gluing agents. As a proof of concept, two novel molecules were described with satisfying biological activity.
In other species
In most of the cell types of most species, most Ras is the GDP type. This is true for Xenopus oocytes and mouse fibroblasts.[33]
Xenopus laevis
As mentioned above most X. oocyte Ras is the GDP conjugate. Mammal Ras induces
Drosophila melanogaster
Expressed in all tissues of Drosophila melanogaster but mostly in neural cells. Overexpression is somewhat lethal and, during development, produces eye and wing abnormalities. (This parallels - and may be the reason for - similar abnormalities due to mutated receptor tyrosine kinases.) The D. genes for rases in mammals produce abnormalities.[33]
Aplysia
Most expression in Aplysia spp. is in neural cells.[33]
Caenorhabditis elegans
The gene in C. elegans is let 60. Also appears to play a role in receptor tyrosine kinase formation in this model. Overexpression yields a multivulval development due to its involvement in that region's normal development; overexpression in effector sites in lethal.[33]
Dictyostelium discoideum
Essential in
References
- PMID 10394594.
- ^ S2CID 43074411.
- ^ S2CID 27928171.
- PMID 6289320.
- S2CID 4144311.
- PMID 5538357.
- S2CID 5807661.
- S2CID 37033023.
- S2CID 4338225.
- S2CID 34179063.
- S2CID 4342747.
- S2CID 4372475.
- PMID 6300838.
- ^ S2CID 6636339.
- PMID 35536216. Retrieved 11 October 2022.
- ^ ISBN 0-7167-3706-X.
- PMID 16781855.
- ^ PMID 2547513.
- S2CID 40171018.
- ^ Robbins and Cotran (2010). Pathologic Basis of Disease 8th ed. p. 282.
- PMID 18374183.
- S2CID 21694752.
- PMID 10961860.
- ^ "Omim - Neuroblastoma Ras Viral Oncogene Homolog; Nras". Archived from the original on 2019-03-06. Retrieved 2017-09-10.
- PMID 19806501.
- PMID 16569513.
- PMID 19565924.
- ^ "ClinicalTrials.gov".
- PMID 26617336.
- ^ Taub B (2015-12-03). "Scientists Find Way To Deactivate Most Common Cancer-Causing Protein". IFLScience. Retrieved 2016-02-20.
- S2CID 207494086.
- PMID 38473821.
- ^ PMID 1667084.
Further reading
- Agrawal AG, Somani RR (June 2009). "Farnesyltransferase inhibitor as anticancer agent". Mini Reviews in Medicinal Chemistry. 9 (6): 638–52. PMID 19519490.
External links
- "Brain tumour findings offer hope of new strategy Canadian Cancer Society says" at ncic.cancer.ca
- "Novel cancer treatment gets NCI support" at arstechnica.com
- ras+Proteins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- ras+Genes at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Drosophila Ras oncogene at 85D - The Interactive Fly
- "Animation of ras activation by EGFR"
- "Rascore: A tool for analyzing RAS protein structures"
