O-GlcNAc

O-GlcNAc (short for O-linked GlcNAc or O-linked β-N-acetylglucosamine) is a reversible enzymatic post-translational modification that is found on serine and threonine residues of nucleocytoplasmic proteins. The modification is characterized by a β-glycosidic bond between the hydroxyl group of serine or threonine side chains and N-acetylglucosamine (GlcNAc). O-GlcNAc differs from other forms of protein glycosylation: (i) O-GlcNAc is not elongated or modified to form more complex glycan structures, (ii) O-GlcNAc is almost exclusively found on nuclear and cytoplasmic proteins rather than membrane proteins and secretory proteins, and (iii) O-GlcNAc is a highly dynamic modification that turns over more rapidly than the proteins which it modifies. O-GlcNAc is conserved across metazoans.[1]

Due to the dynamic nature of O-GlcNAc and its presence on serine and threonine residues, O-GlcNAcylation is similar to protein phosphorylation in some respects. While there are roughly 500 kinases and 150 phosphatases that regulate protein phosphorylation in humans, there are only 2 enzymes that regulate the cycling of O-GlcNAc: O-GlcNAc transferase (OGT) and O-GlcNAcase (OGA) catalyze the addition and removal of O-GlcNAc, respectively.[2] OGT utilizes UDP-GlcNAc as the donor sugar for sugar transfer.[3]
First reported in 1984, this post-translational modification has since been identified on over 5,000 proteins.
Discovery

In 1984, the Hart lab was probing for terminal GlcNAc residues on the surfaces of thymocytes and lymphocytes. Bovine milk β-1,4-galactosyltransferase, which reacts with terminal GlcNAc residues, was used to perform radiolabeling with UDP-[3H]galactose. β-elimination of serine and threonine residues demonstrated that most of the [3H]galactose was attached to proteins O-glycosidically; chromatography revealed that the major β-elimination product was Galβ1-4GlcNAcitol. Insensitivity to peptide N-glycosidase treatment provided additional evidence for O-linked GlcNAc. Permeabilizing cells with detergent prior to radiolabeling greatly increased the amount of [3H]galactose incorporated into Galβ1-4GlcNAcitol, leading the authors to conclude that most of the O-linked GlcNAc monosaccharide residues were intracellular.[12]
Mechanism

O-GlcNAc is generally a dynamic modification that can be cycled on and off various proteins. Some residues are thought to be constitutively modified by O-GlcNAc.[13][14] The O-GlcNAc modification is installed by OGT in a sequential bi-bi mechanism where the donor sugar, UDP-GlcNAc, binds to OGT first followed by the substrate protein.[15] The O-GlcNAc modification is removed by OGA in a hydrolysis mechanism involving anchimeric assistance (substrate-assisted catalysis) to yield the unmodified protein and GlcNAc.[16] While crystal structures have been reported for both OGT[15] and OGA,[17][18] the exact mechanisms by which OGT and OGA recognize substrates have not been completely elucidated. Unlike N-linked glycosylation, for which glycosylation occurs in a specific consensus sequence (Asn-X-Ser/Thr, where X is any amino acid except Pro), no definitive consensus sequence has been identified for O-GlcNAc,. Consequently, predicting sites of O-GlcNAc modification is challenging, and identifying modification sites generally requires mass spectrometry methods. For OGT, studies have shown that substrate recognition is regulated by a number of factors including aspartate[19] and asparagine[20] ladder motifs in the lumen of the superhelical TPR domain, active site residues,[21] and adaptor proteins.[22] As crystal structures have shown that OGT requires its substrate to be in an extended conformation, it has been proposed that OGT has a preference for flexible substrates.[21] In in vitro kinetic experiments measuring OGT and OGA activity on a panel of protein substrates, kinetic parameters for OGT were shown to be variable between various proteins while kinetic parameters for OGA were relatively constant between various proteins. This result suggested that OGT is the "senior partner" in regulating O-GlcNAc and OGA primarily recognizes substrates via the presence of O-GlcNAc rather than the identity of the modified protein.[13]

Detection and characterization
Several methods exist to detect the presence of O-GlcNAc and characterize the specific residues modified.
Lectins
Wheat germ agglutinin, a plant lectin, is able to recognize terminal GlcNAc residues and is thus often used for detection of O-GlcNAc. This lectin has been applied in lectin affinity chromatography for the enrichment and detection of O-GlcNAc.[23]
Antibodies
Pan-O-GlcNAc antibodies that recognize the O-GlcNAc modification largely irrespective of the modified protein's identity are commonly used. These include RL2,[24] an IgG antibody raised against O-GlcNAcylated nuclear pore complex proteins, and CTD110.6,[25] an IgM antibody raised against an immunogenic peptide with a single serine O-GlcNAc modification. Other O-GlcNAc-specific antibodies have been reported and demonstrated to have some dependence on the identity of the modified protein.[26]
Metabolic labeling
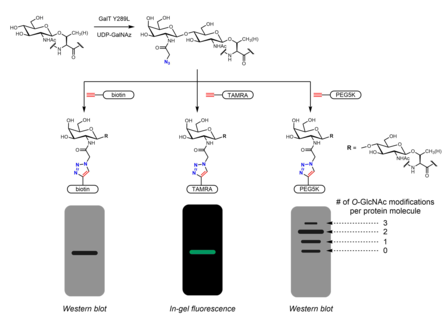
Many metabolic chemical reporters have been developed to identify O-GlcNAc. Metabolic chemical reporters are generally sugar analogues that bear an additional chemical moiety allowing for additional reactivity. For example, peracetylated GlcNAc (Ac4GlcNAz) is a cell-permeable azido sugar that is de-esterified intracellularly by esterases to GlcNAz and converted to UDP-GlcNAz in the hexosamine salvage pathway. UDP-GlcNAz can be utilized as a sugar donor by OGT to yield the O-GlcNAz modification.[27] The presence of the azido sugar can then be visualized via alkyne-containing bioorthogonal chemical probes in an azide-alkyne cycloaddition reaction. These probes can incorporate easily identifiable tags such as the FLAG peptide, biotin, and dye molecules.[27][28] Mass tags based on polyethylene glycol (PEG) have also been used to measure O-GlcNAc stoichiometry. Conjugation of 5 kDa PEG molecules leads to a mass shift for modified proteins - more heavily O-GlcNAcylated proteins will have multiple PEG molecules and thus migrate more slowly in gel electrophoresis.[29] Other metabolic chemical reporters bearing azides or alkynes (generally at the 2 or 6 positions) have been reported.[30] Instead of GlcNAc analogues, GalNAc analogues may be used as well as UDP-GalNAc is in equilibrium with UDP-GlcNAc in cells due to the action of UDP-galactose-4'-epimerase (GALE). Treatment with Ac4GalNAz was found to result in enhanced labeling of O-GlcNAc relative to Ac4GlcNAz, possibly due to a bottleneck in UDP-GlcNAc pyrophosphorylase processing of GlcNAz-1-P to UDP-GlcNAz.[31] Ac3GlcN-β-Ala-NBD-α-1-P(Ac-SATE)2, a metabolic chemical reporter that is processed intracellularly to a fluorophore-labeled UDP-GlcNAc analogue, has been shown to achieve one-step fluorescent labeling of O-GlcNAc in live cells.[32]
Metabolic labeling may also be used to identify binding partners of O-GlcNAcylated proteins. The N-acetyl group may be elongated to incorporate a diazirine moiety. Treatment of cells with peracetylated, phosphate-protected Ac3GlcNDAz-1-P(Ac-SATE)2 leads to modification of proteins with O-GlcNDAz. UV irradiation then induces photocrosslinking between proteins bearing the O-GlcNDaz modification and interacting proteins.[33]
Some issues have been identified with various metabolic chemical reporters, e.g., their use may inhibit the hexosamine biosynthetic pathway,[30] they may not be recognized by OGA and therefore are not able to capture O-GlcNAc cycling,[34] or they may be incorporated into glycosylation modifications besides O-GlcNAc as seen in secreted proteins.[35] Metabolic chemical reporters with chemical handles at the N-acetyl position may also label acetylated proteins as the acetyl group may be hydrolyzed into acetate analogues that can be utilized for protein acetylation.[36] Additionally, per-O-acetylated monosaccharides have been identified to react cysteines leading to artificial S-glycosylation.[37] This occurs via an elimination-addition mechanism.[38]
Chemoenzymatic labeling
Chemoenzymatic labeling provides an alternative strategy to incorporate handles for click chemistry. The Click-IT O-GlcNAc Enzymatic Labeling System, developed by the Hsieh-Wilson group and subsequently commercialized by Invitrogen, utilizes a mutant GalT Y289L enzyme that is able to transfer azidogalactose (GalNAz) onto O-GlcNAc.[28][39] The presence of GalNAz (and therefore also O-GlcNAc) can be detected with various alkyne-containing probes with identifiable tags such as biotin,[39] dye molecules,[28] and PEG.[29]

Förster resonance energy transfer biosensor
An engineered protein biosensor has been developed that can detect changes in O-GlcNAc levels using Förster resonance energy transfer. This sensor consists of four components linked together in the following order: cyan fluorescent protein (CFP), an O-GlcNAc binding domain (based on GafD, a lectin sensitive for terminal β-O-GlcNAc), a CKII peptide that is a known OGT substrate, and yellow fluorescent protein (YFP). Upon O-GlcNAcylation of the CKII peptide, the GafD domain binds the O-GlcNAc moiety, bringing the CFP and YFP domains into close proximity and generating a FRET signal. Generation of this signal is reversible and can be used to monitor O-GlcNAc dynamics in response to various treatments. This sensor may be genetically encoded and used in cells.[40] Addition of a localization sequence allows for targeting of this O-GlcNAc sensor to the nucleus, cytoplasm, or plasma membrane.[41]
Mass spectrometry
Biochemical approaches such as Western blotting may provide supporting evidence that a protein is modified by O-GlcNAc; mass spectrometry (MS) is able to provide definitive evidence as to the presence of O-GlcNAc. Glycoproteomic studies applying MS have contributed to the identification of proteins modified by O-GlcNAc.
As O-GlcNAc is substoichiometric and
Traditional proteomic studies perform tandem MS on the most abundant species in the full-scan mass spectra, prohibiting full characterization of lower-abundance species. One modern strategy for targeted proteomics uses isotopic labels, e.g., dibromide, to tag O-GlcNAcylated proteins. This method allows for algorithmic detection of low-abundance species, which are then sequenced by tandem MS.[45] Directed tandem MS and targeted glycopeptide assignment allow for identification of O-GlcNAcylated peptide sequences. One example probe consists of a biotin affinity tag, an acid-cleavable silane, an isotopic recoding motif, and an alkyne.[46][47][48] Unambiguous site mapping is possible for peptides with only one serine/threonine residue.[49]
The general procedure for this isotope-targeted glycoproteomics (IsoTaG) method is the following:

- Metabolically label O-GlcNAc to install O-GlcNAz onto proteins
- Use click chemistry to link IsoTaG probe to O-GlcNAz
- Use streptavidin beads to enrich for tagged proteins
- Treat beads with trypsin to release non-modified peptides
- Cleave isotopically recoded glycopeptides from beads using mild acid
- Obtain a full-scan mass spectrum from isotopically recoded glycopeptides
- Apply algorithm to detect unique isotope signature from probe
- Perform tandem MS on the isotopically recoded species to obtain glycopeptide amino acid sequences
- Search protein database for identified sequences
Other methodologies have been developed for quantitative profiling of O-GlcNAc using differential isotopic labeling.[50] Example probes generally consist of a biotin affinity tag, a cleavable linker (acid- or photo-cleavable), a heavy or light isotopic tag, and an alkyne.[51][52]
Strategies for manipulating O-GlcNAc
Various chemical and genetic strategies have been developed to manipulate O-GlcNAc, both on a proteome-wide basis and on specific proteins.
Chemical methods
Small molecule inhibitors have been reported for both OGT[53][54] and OGA[55][56] that function in cells or in vivo. OGT inhibitors result in a global decrease of O-GlcNAc while OGA inhibitors result in a global increase of O-GlcNAc; these inhibitors are not able to modulate O-GlcNAc on specific proteins.
Inhibition of the hexosamine biosynthetic pathway is also able to decrease O-GlcNAc levels. For instance, glutamine analogues azaserine and 6-diazo-5-oxo-L-norleucine (DON) can inhibit GFAT, though these molecules may also non-specifically affect other pathways.[57]
Protein synthesis
Genetic methods
Site-directed mutagenesis

Site-directed mutagenesis of O-GlcNAc-modified serine or threonine residues to alanine may be used to evaluate the function of O-GlcNAc at specific residues. As alanine's side chain is a methyl group and is thus not able to act as an O-GlcNAc site, this mutation effectively permanently removes O-GlcNAc at a specific residue. While serine/threonine phosphorylation may be modeled by mutagenesis to aspartate or glutamate, which have negatively charged carboxylate side chains, none of the 20 canonical amino acids sufficiently recapitulate the properties of O-GlcNAc.[60] Mutagenesis to tryptophan has been used to mimic the steric bulk of O-GlcNAc, though tryptophan is much more hydrophobic than O-GlcNAc.[61][62] Mutagenesis may also perturb other post-translational modifications, e.g., if a serine is alternatively phosphorylated or O-GlcNAcylated, alanine mutagenesis permanently eliminates the possibilities of both phosphorylation and O-GlcNAcylation.
S-GlcNAc
Mass spectrometry identified S-GlcNAc as a post-translational modification found on cysteine residues. In vitro experiments demonstrated that OGT could catalyze the formation of S-GlcNAc and that OGA is incapable of hydrolyzing S-GlcNAc.[63] Though a previous report suggested that OGA is capable of hydrolyzing thioglycosides, this was only demonstrated on the aryl thioglycoside para-nitrophenol-S-GlcNAc; para-nitrothiophenol is a more activated leaving group than a cysteine residue.[64] Recent studies have supported the use of S-GlcNAc as an enzymatically stable structural model of O-GlcNAc that can be incorporated through solid-phase peptide synthesis or site-directed mutagenesis.[65][60][58][66]
Engineered OGT
Fusion constructs of a nanobody and TPR-truncated OGT allow for proximity-induced protein-specific O-GlcNAcylation in cells. The nanobody may be directed towards protein tags, e.g., GFP, that are fused to the target protein, or the nanobody may be directed towards endogenous proteins. For example, a nanobody recognizing a C-terminal EPEA sequence can direct OGT enzymatic activity to α-synuclein.[67]
Functions of O-GlcNAc
Apoptosis
Apoptosis, a form of controlled cell death, has been suggested to be regulated by O-GlcNAc. In various cancers, elevated O-GlcNAc levels have been reported to suppress apoptosis.[68][69] Caspase-3, caspase-8, and caspase-9 have been reported to be modified by O-GlcNAc. Caspase-8 is modified near its cleavage/activation sites; O-GlcNAc modification may block caspase-8 cleavage and activation by steric hindrance. Pharmacological lowering of O-GlcNAc with 5S-GlcNAc accelerated caspase activation while pharmacological raising of O-GlcNAc with thiamet-G inhibited caspase activation.[62]
Epigenetics
Writers and Erasers
The proteins that regulate genetics are often categorized as writers, readers, and erasers, i.e., enzymes that install epigenetic modifications, proteins that recognize these modifications, and enzymes that remove these modifications.[70] To date, O-GlcNAc has been identified on writer and eraser enzymes. O-GlcNAc is found in multiple locations on EZH2, the catalytic methyltransferase subunit of PRC2, and is thought to stabilize EZH2 prior to PRC2 complex formation and regulate di- and tri-methyltransferase activity.[71][72] All three members of the ten-eleven translocation (TET) family of dioxygenases (TET1, TET2, and TET3) are known to be modified by O-GlcNAc.[73] O-GlcNAc has been suggested to cause nuclear export of TET3, reducing its enzymatic activity by depleting it from the nucleus.[74] O-GlcNAcylation of HDAC1 is associated with elevated activating phosphorylation of HDAC1.[75]
Histone O-GlcNAcylation
Histone proteins, the primary protein component of chromatin, are known to be modified by O-GlcNAc.[8] O-GlcNAc has been identified on all core histones (H2A,[8] H2B,[8] H3,[76] and H4[8]). The presence of O-GlcNAc on histones has been suggested to affect gene transcription as well as other histone marks such as acetylation[8] and monoubiquitination.[77] TET2 has been reported to interact with the TPR domain of OGT and facilitate recruitment of OGT to histones.[78] This interaction is associated with H2B S112 O-GlcNAc, which in turn is associated with H2B K120 monoubiquitination.[77] Phosphorylation of OGT T444 via AMPK has been found to inhibit OGT-chromatin association and downregulate H2B S112 O-GlcNAc.[79]
Nutrient sensing
The hexosamine biosynthetic pathway's product, UDP-GlcNAc, is utilized by OGT to catalyze the addition of O-GlcNAc. This pathway integrates information about the concentrations of various metabolites including amino acids, carbohydrates, fatty acids, and nucleotides. Consequently, UDP-GlcNAc levels are sensitive to cellular metabolite levels. OGT activity is in part regulated by UDP-GlcNAc concentration, making a link between cellular nutrient status and O-GlcNAc.[80]
Glucose deprivation causes a decline in UDP-GlcNAc levels and an initial decline in O-GlcNAc, but counterintuitively, O-GlcNAc is later significantly upregulated. This later increase has been shown to be dependent on AMPK and p38 MAPK activation, and this effect is partially due to increases in OGT mRNA and protein levels.[81] It has also been suggested that this effect is dependent on calcium and CaMKII.[82] Activated p38 is able to recruit OGT to specific protein targets, including neurofilament H; O-GlcNAc modification of neurofilament H enhances its solubility.[81] During glucose deprivation, glycogen synthase is modified by O-GlcNAc which inhibits its activity.[83]
Oxidative stress
NRF2, a transcription factor associated with the cellular response to oxidative stress, has been found to be indirectly regulated by O-GlcNAc. KEAP1, an adaptor protein for the cullin 3-dependent E3 ubiquitin ligase complex, mediates the degradation of NRF2; oxidative stress leads to conformational changes in KEAP1 that repress degradation of NRF2. O-GlcNAc modification of KEAP1 at S104 is required for efficient ubiquitination and subsequent degradation of NRF2, linking O-GlcNAc to oxidative stress. Glucose deprivation leads to a reduction in O-GlcNAc and reduces NRF2 degradation. Cells expressing a KEAP1 S104A mutant are resistant to erastin-induced ferroptosis, consistent with higher NRF2 levels upon removal of S104 O-GlcNAc.[84]
Elevated O-GlcNAc levels have been associated with diminished synthesis of hepatic glutathione, an important cellular antioxidant. Acetaminophen overdose leads to accumulation of the strongly oxidizing metabolite NAPQI in the liver, which is detoxified by glutathione. In mice, OGT knockout has a protective effect against acetaminophen-induced liver injury, while OGA inhibition with thiamet-G exacerbates acetaminophen-induced liver injury.[85]
Protein aggregation
O-GlcNAc has been found to slow protein aggregation, though the generality of this phenomenon is unknown.
Solid-phase peptide synthesis was used to prepare full-length α-synuclein with an O-GlcNAc modification at T72. Thioflavin T aggregation assays and transmission electron microscopy demonstrated that this modified α-synuclein does not readily form aggregates.[59]
Treatment of JNPL3 tau transgenic mice with an OGA inhibitor was shown to increase microtubule-associated protein tau O-GlcNAcylation. Immunohistochemistry analysis of the brainstem revealed decreased formation of neurofibrillary tangles. Recombinant O-GlcNAcylated tau was shown to aggregate slower than unmodified tau in an in vitro thioflavin S aggregation assay. Similar results were obtained for a recombinantly prepared O-GlcNAcylated TAB1 construct versus its unmodified form.[86]
Protein phosphorylation
Crosstalk
Many known phosphorylation sites and O-GlcNAcylation sites are nearby each other or overlapping.
In some cases, therapeutic strategies are under investigation to modulate O-GlcNAcylation to have a downstream effect on phosphorylation. For instance, elevating tau O-GlcNAcylation may offer therapeutic benefit by inhibiting pathological tau hyperphosphorylation.[95]
Besides phosphorylation, O-GlcNAc has been found to influence other post-translational modifications such as lysine acetylation[92] and monoubiquitination.[77]
Kinases
Protein kinases are the enzymes responsible for phosphorylation of serine and threonine residues. O-GlcNAc has been identified on over 100 (~20% of the human kinome) kinases, and this modification is often associated with alterations in kinase activity or kinase substrate scope.[96]
The first report of a kinase being directly regulated by O-GlcNAc was published in 2009. CaMKIV is glycosylated at multiple sites, though S189 was found to be the major site. An S189A mutant was more readily activated by CaMKIV T200 phosphorylation, suggesting that O-GlcNAc at S189 inhibits CaMKIV activity. Homology modeling showed that S189 O-GlcNAc may interfere with ATP binding.[91]
AMPK and OGT are known to modify each other, i.e., AMPK phosphorylates OGT and OGT O-GlcNAcylates AMPK. AMPK activation by AICA ribonucleotide is associated with nuclear localization of OGT in differentiated C2C12 mouse skeletal muscle myotubes, resulting in increased nuclear O-GlcNAc. This effect was not observed in proliferating cells and undifferentiated myoblastic cells.[97] AMPK phosphorylation of OGT T444 has been found to block OGT association with chromatin and decrease H2B S112 O-GlcNAc.[79] Overexpression of GFAT, the enzyme that controls glucose flux into the hexosamine biosynthetic pathway, in mouse adipose tissue has been found to lead to AMPK activation and downstream ACC inhibition and elevated fatty acid oxidation. Glucosamine treatment in cultured 3T3L1 adipocytes showed a similar effect.[98] The exact relationship between O-GlcNAc and AMPK has not been completely elucidated as various studies have reported that OGA inhibition inhibits AMPK activation,[97] OGT inhibition also inhibits AMPK activation,[79] upregulating O-GlcNAc by glucosamine treatment activates AMPK,[98] and OGT knockdown activates AMPK;[99] these results suggest that additional indirect communication between AMPK pathways and O-GlcNAc or cell type-specific effects.
CKIIα substrate recognition has been shown to be altered upon S347 O-GlcNAcylation.[58]
Phosphatases
Protein phosphatase 1 subunits PP1β and PP1γ have been shown to form functional complexes with OGT. A synthetic phosphopeptide was able to be dephosphorylated and O-GlcNAcylated by an OGT immunoprecipitate. This complex has been referred to as a "yin-yang complex" as it replaces a phosphate modification with an O-GlcNAc modification.[100]
MYPT1 is another protein phosphatase subunit that forms complexes with OGT and is itself O-GlcNAcylated. MYPT1 appears to have a role in directing OGT towards specific substrates.[101]
Protein-protein interactions
O-GlcNAcylation of a protein can alter its interactome. As O-GlcNAc is highly hydrophilic, its presence may disrupt hydrophobic protein-protein interactions. For example, O-GlcNAc disrupts
Some studies have also identified instances where protein-protein interactions are induced by O-GlcNAc. Metabolic labeling with the diazirine-containing O-GlcNDAz has been applied to identify protein-protein interactions induced by O-GlcNAc.
Protein stability and degradation
Co-translational O-GlcNAc has been identified on
Protein phosphorylation is often used as a mark for subsequent degradation.
O-GlcNAcylation of the Rpt2 ATPase subunit of the 26S proteasome has been shown to inhibit proteasome activity. Testing various peptide sequences revealed that this modification slows proteasomal degradation of hydrophobic peptides, degradation of hydrophilic peptides does not appear to be affected.[107] This modification has been shown to suppress other pathways that activate the proteasome such as Rpt6 phosphorylation by cAMP-dependent protein kinase.[108]
OGA-S localizes to lipid droplets and has been proposed to locally activate the proteasome to promote remodeling of lipid droplet surface proteins.[109]
Stress response
Various cellular stress stimuli have been associated with changes in O-GlcNAc. Treatment with hydrogen peroxide, cobalt(II) chloride, UVB light, ethanol, sodium chloride, heat shock, and sodium arsenite, all result in elevated O-GlcNAc. Knockout of OGT sensitizes cells to thermal stress. Elevated O-GlcNAc has been associated with expression of Hsp40 and Hsp70.[110]
Therapeutic relevance
Alzheimer's disease
Numerous studies have identified aberrant phosphorylation of tau as a hallmark of Alzheimer's disease.[111] O-GlcNAcylation of bovine tau was first characterized in 1996.[112] A subsequent report in 2004 demonstrated that human brain tau is also modified by O-GlcNAc. O-GlcNAcylation of tau was demonstrated to regulate tau phosphorylation with hyperphosphorylation of tau observed in the brain of mice lacking OGT,[113] which has been associated with the formation of neurofibrillary tangles. Analysis of brain samples showed that protein O-GlcNAcylation is compromised in Alzheimer's disease and paired helical fragment-tau was not recognized by traditional O-GlcNAc detection methods, suggesting that pathological tau has impaired O-GlcNAcylation relative to tau isolated from control brain samples. Elevating tau O-GlcNAcylation was proposed as a therapeutic strategy for reducing tau phosphorylation.[89]
To test this therapeutic hypothesis, a selective and blood-brain barrier-permeable OGA inhibitor, thiamet-G, was developed. Thiamet-G treatment was able to increase tau O-GlcNAcylation and suppress tau phosphorylation in cell culture and in vivo in healthy Sprague-Dawley rats.[56] A subsequent study showed that thiamet-G treatment also increased tau O-GlcNAcylation in a JNPL3 tau transgenic mouse model. In this model, tau phosphorylation was not significantly affected by thiamet-G treatment, though decreased numbers of neurofibrillary tangles and slower motor neuron loss were observed. Additionally, O-GlcNAcylation of tau was noted to slow tau aggregation in vitro.[86]
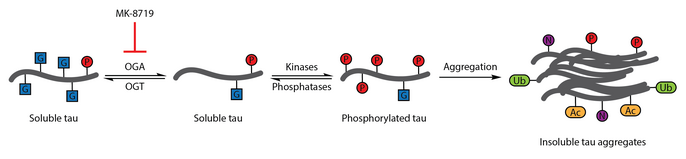
OGA inhibition with MK-8719 is being investigated in clinical trials as a potential treatment strategy for Alzheimer's disease and other tauopathies including progressive supranuclear palsy.[95][114][115]
Cancer
Dysregulation of O-GlcNAc is associated with cancer cell proliferation and tumor growth.
O-GlcNAcylation of the
Human pancreatic ductal adenocarcinoma (PDAC) cell lines have higher O-GlcNAc levels than human pancreatic duct epithelial (HPDE) cells. PDAC cells have some dependency upon O-GlcNAc for survival as OGT knockdown selectively inhibited PDAC cell proliferation (OGT knockdown did not significantly affect HPDE cell proliferation), and inhibition of OGT with 5S-GlcNAc showed the same result. Hyper-O-GlcNAcylation in PDAC cells appeared to be anti-apoptotic, inhibiting cleavage and activation of caspase-3 and caspase-9. Numerous sites on the p65 subunit of NF-κB were found to be modified by O-GlcNAc in a dynamic manner; O-GlcNAc at p65 T305 and S319 in turn positively regulate other modifications associated with NF-κB activation such as p300-mediated K310 acetylation and IKK-mediated S536 phosphorylation. These results suggested that NF-κB is constitutively activated by O-GlcNAc in pancreatic cancer.[69][92]
OGT stabilization of EZH2 in various breast cancer cell lines has been found to inhibit expression of tumor suppressor genes.
OGT has been found to stabilize SREBP-1 and activate lipogenesis in breast cancer cell lines. This stabilization was dependent on the proteasome and AMPK. OGT knockdown resulted in decreased nuclear SREBP-1, but proteasomal inhibition with MG132 blocked this effect. OGT knockdown also increased the interaction between SREBP-1 and the E3 ubiquitin ligase FBW7. AMPK is activated by T172 phosphorylation upon OGT knockdown, and AMPK phosphorylates SREBP-1 S372 to inhibit its cleavage and maturation. OGT knockdown had a diminished effect on SREBP-1 levels in AMPK-null cell lines. In a mouse model, OGT knockdown inhibited tumor growth but SREBP-1 overexpression partly rescued this effect.[99] These results contrast from those of a previous study which found that OGT knockdown/inhibition inhibited AMPK T172 phosphorylation and increased lipogenesis.[79]
In breast and prostate cancer cell lines, high levels of OGT and O-GlcNAc have been associated both in vitro and in vivo with processes associated with disease progression, e.g.,
Diabetes
Elevated O-GlcNAc has been associated with diabetes.
Pancreatic β cells synthesize and secrete insulin to regulate blood glucose levels. One study found that inhibition of OGA with streptozotocin followed by glucosamine treatment resulted in O-GlcNAc accumulation and apoptosis in β cells;[119] a subsequent study showed that a galactose-based analogue of streptozotocin was unable to inhibit OGA but still resulted in apoptosis, suggesting that the apoptotic effects of streptozotocin are not directly due to OGA inhibition.[120]
O-GlcNAc has been suggested to attenuate
As PUGNAc also inhibits lysosomal β-hexosaminidases, the OGA-selective inhibitor NButGT was developed to further probe the relationship between O-GlcNAc and insulin signaling in 3T3-L1 adipocytes. This study also found that PUGNAc resulted in impaired insulin signaling, but NButGT did not, as measured by changes in phosphorylation of Akt T308, suggesting that the effects observed with PUGNAc may be due to off-target effects besides OGA inhibition.[124]
Parkinson's disease
Parkinson's disease is associated with aggregation of α-synuclein.[125] As O-GlcNAc modification of α-synuclein has been found to inhibit its aggregation, elevating α-synuclein O-GlcNAc is being explored as a therapeutic strategy to treat Parkinson's disease.[59][126]
Infectious disease
Bacterial
Treatment of macrophages with lipopolysaccharide (LPS), a major component of the Gram-negative bacteria outer membrane, results in elevated O-GlcNAc in cellular and mouse models. During infection, cytosolic OGT was de-S-nitrosylated and activated. Suppressing O-GlcNAc with DON inhibited the O-GlcNAcylation and nuclear translocation of NF-κB, as well as downstream induction of inducible nitric oxide synthase and IL-1β production. DON treatment also improved cell survival during LPS treatment.[127]
Viral
O-GlcNAc has been implicated in influenza A virus (IAV)-induced cytokine storm. Specifically, O-GlcNAcylation of S430 on interferon regulatory factor-5 (IRF5) has been shown to promote its interaction with TNF receptor-associated factor 6 (TRAF6) in cellular and mouse models. TRAF6 mediates K63-linked ubiquitination of IRF5 which is necessary for IRF5 activity and subsequent cytokine production. Analysis of clinical samples showed that blood glucose levels were elevated in IAV-infected patients compared to healthy individuals. In IAV-infected patients, blood glucose levels positively correlated with IL-6 and IL-8 levels. O-GlcNAcylation of IRF5 was also relatively higher in peripheral blood mononuclear cells of IAV-infected patients.[128]
Other applications
Peptide therapeutics such as are attractive for their high specificity and potency, but they often have poor pharmacokinetic profiles due to their degradation by serum proteases.[129] Though O-GlcNAc is generally associated with intracellular proteins, it has been found that engineered peptide therapeutics modified by O-GlcNAc have enhanced serum stability in a mouse model and have similar structure and activity compared to the respective unmodified peptides. This method has been applied to engineer GLP-1 and PTH peptides.[130]
See also
References
- ^ PMID 20016062.
- PMID 22564745.
- PMID 2137449.
- PMID 33479245.
- PMID 24593906.
- PMID 35230102.
- PMID 8486697.
- ^ PMID 21045127.
- PMID 27294441.
- PMID 30464755.
- PMID 21391816.
- PMID 6421821.
- ^ PMID 22311971.
- ^ PMID 25774941.
- ^ PMID 21240259.
- PMID 15795231.
- PMID 28346405.
- PMID 28346407.
- PMID 31373491.
- PMID 29485866.
- ^ PMID 26237509.
- PMID 18840611.
- PMID 22045558.
- PMID 2437126.
- PMID 11399029.
- PMID 20305658.
- ^ PMID 12874386.
- ^ PMID 18683930.
- ^ PMID 20657584.
- ^ PMID 29979493.
- PMID 21300897.
- S2CID 207194442.
- ^ PMID 22411826.
- PMID 25068034.
- PMID 21540332.
- PMID 25062036.
- PMID 29237092.
- PMID 32339456.
- ^ a b "Click-IT™ O-GlcNAc Enzymatic Labeling System". www.thermofisher.com. Retrieved 2020-05-30.
- PMID 17105262.
- PMID 21138847.
- PMID 28150883.
- PMID 12438562.
- PMID 21740066.
- PMID 21604797.
- PMID 25894945.
- PMID 28244757.
- PMID 27695962.
- ^ PMID 29351928.
- PMID 17496889.
- S2CID 206528204.
- S2CID 58620368.
- PMID 29756380.
- PMID 30285435.
- PMID 17177381.
- ^ PMID 18587388.
- PMID 31272438.
- ^ PMID 22267120.
- ^ PMID 26492012.
- ^ PMID 31695185.
- PMID 28195695.
- ^ PMID 28528544.
- PMID 27558639.
- PMID 16332065.
- PMID 32610041.
- PMID 28627871.
- PMID 32119511.
- ^ PMID 24857547.
- ^ PMID 23592772.
- PMID 26496625.
- ^ PMID 24474760.
- PMID 29941599.
- PMID 24394411.
- PMID 24394411.
- ^ PMID 27060025.
- PMID 22371497.
- ^ PMID 22121020.
- PMID 23222540.
- ^ PMID 24692660.
- PMID 10542233.
- ^ PMID 18353774.
- PMID 22908225.
- PMID 18174169.
- PMID 28663241.
- PMID 29325178.
- ^ PMID 22366723.
- PMID 10995228.
- PMID 11425311.
- ^ PMID 15249677.
- ^ S2CID 12326082.
- ^ PMID 19506079.
- ^ PMID 28416608.
- ^ PMID 24744147.
- ^ PMID 24214978.
- ^ PMID 31487175.
- PMID 32155042.
- ^ PMID 24563466.
- ^ PMID 17227772.
- ^ PMID 29059153.
- PMID 15247246.
- PMID 18840611.
- PMID 11371615.
- PMID 12769553.
- PMID 22267118.
- PMID 29784830.
- PMID 15158436.
- S2CID 8221476.
- PMID 17565987.
- PMID 21807949.
- PMID 15138254.
- PMID 20678074.
- PMID 8910513.
- ^ O'Donnell, N, Zachara, N, Hart, GW, Marth JD. (2004). Ogt-dependent X-chromosome-linked protein glycosylation is a requisite modification in somatic cell function and embryo viability. Molecular and Cellular Biology. 24: 1680-1690. Diol.
- S2CID 54229492.
- PMID 29641484.
- PMID 22923583.
- S2CID 25957261.
- PMID 22275356.
- PMID 10717000.
- PMID 18721751.
- ^ PMID 11959983.
- S2CID 18459576.)
{{cite journal}}: CS1 maint: numeric names: authors list (link - PMID 18519567.
- PMID 18842583.
- PMID 22355802.
- ^ "Glycosylation as an Inhibitor of Alpha-synuclein Aggregation". The Michael J. Fox Foundation for Parkinson's Research | Parkinson's Disease. Retrieved 2020-06-05.
- PMID 21453677.
- PMID 32494619.
- PMID 23726889.
- PMID 31418572.
Further reading
- Zachara, Natasha; Akimoto, Yoshihiro; Hart, Gerald W. (2015), Varki, Ajit; Cummings, Richard D.; Esko, Jeffrey D.; Stanley, Pamela (eds.), "The O-GlcNAc Modification", Essentials of Glycobiology (3rd ed.), Cold Spring Harbor Laboratory Press, PMID 28876858.
