Evolution of the wolf

The evolution of the wolf occurred over a geologic time scale of at least 300 thousand years. The
The archaeological and paleontological records show the grey wolf's continuous presence for at least the last 300,000 years.[8] This continuous presence contrasts with genomic analyses, which suggest that all modern wolves and dogs descend from a common ancestral wolf population[9][10][11] that existed as recently as 20,000 years ago.[9] Grey wolves suffered a species-wide population bottleneck (reduction) approximately 25,000 YBP during the Last Glacial Maximum. This was followed by a single population of modern wolves expanding out of a Beringia refuge to repopulate the wolf's former range, replacing the remaining Late Pleistocene wolf populations across Eurasia and North America as they did so.[12]
Fossil record

The fossil record for ancient vertebrates is composed of rarely occurring fragments from which it is often impossible to obtain genetic material. Researchers are limited to morphologic analysis but it is difficult to estimate the intra-species and inter-species variations and relationships that existed between specimens across time and place. Some observations are debated by researchers who do not always agree, and hypotheses that are supported by some authors are challenged by others.[13]
The
The carnivoran ancestors of the dog-like
The canids that had immigrated from North America to Eurasia –
The fossil record is incomplete but it is likely that wolves arose from a population of small, early canids.[23]: p241 Morphological evidence[23]: p239 [24] and genetic evidence[25] both suggest that wolves evolved during the Pliocene and Early Pleistocene eras from the same lineage that also led to the coyote,[23]: p239 with fossil specimens indicating that the coyote and the wolf diverged from a common ancestor 1.5 million years ago.[23]: p240 [24] The ancestor of the jackal and the other extant members of the genus Canis had split from the lineage before this time.[23]: p240
After this separation from a common ancestor the species that were believed to be involved in the further evolution of the wolf and coyote – and the beliefs of some
| Wolf evolution | |||||||||||||||||||||||||||
| |||||||||||||||||||||||||||
| Proposed evolution and branching of genus Canis towards the wolf.[23]: p240 |
Canis lepophagus
Canis lepophagus lived in the early Pliocene in North America.[32] Kurten proposed that the Blancan C. lepophagus[33] derived from smaller Miocene Canis species in North America. It then became widespread across Eurasia where it was either identical to, or closely related with, C. arnensis of Europe.[23]: p241 [34][35]
Johnston describes C. lepophagus as having a more slender skull and skeleton than in the modern coyote.[36]: 385 Robert M. Nowak found that the early populations had small, delicate and narrowly proportioned skulls that resemble small coyotes and appear to be ancestral to C. latrans.[23]: p241 Johnson noted that some specimens found in Cita Canyon, Texas, had larger, broader skulls,[36] and along with other fragments Nowak suggested that these were evolving into wolves.[23]: p241 [24]
Tedford disagreed with previous authors and found that its cranio-dental morphology lacked some characteristics that are shared by C. lupus and C. latrans, and therefore there was not a close relationship but it did suggest C. lepophagus was the ancestor of both wolves and coyotes.[37]: p119
Canis armbrusteri
The North American wolves became larger, with tooth specimens indicating that C. priscolatrans diverged into the large wolf
Aenocyon dirus
In 1908 the paleontologist John Campbell Merriam began retrieving numerous fossilized bone fragments of a large wolf from the Rancho La Brea tar pits. By 1912 he had found a skeleton sufficiently complete to be able to formally recognize these and the previously found specimens under the name C. dirus (Leidy 1858)[citation needed].
Canis dirus,
However, in 2021, a study indicated the dire wolf to be a highly divergent lineage which last shared a most recent common ancestor with the wolf-like canines 5.7 million years ago. The morphological similarity between dire wolves and gray wolves was concluded to be due to convergent evolution. This finding indicates that the wolf and coyote lineages evolved in isolation from the dire wolf lineage. The study proposes an early origin of the dire wolf lineage in the Americas, and that this geographic isolation allowed them to develop a degree of reproductive isolation since their divergence 5.7 million years ago. Coyotes, dholes, gray wolves, and the extinct Xenocyon evolved in Eurasia and expanded into North America relatively recently during the Late Pleistocene, therefore there was no admixture with the dire wolf. As a result, the study found that the correct binomial name of the dire wolf is Aenocyon dirus, as proposed by Merriam in 1918. The long-term isolation of the dire wolf lineage implies that other American fossil taxa, including C. armbrusteri and C. edwardii, may also belong to the dire wolf's lineage.[48]
dirus–lupus hybrids
Merriam named 3 unusual species based on specimens recovered from the Rancho
Canis occidentalis furlongi (Merriam 1910)
Canis milleri (Merriam 1912),[51] the Miller wolf, was as large as the timber wolf but with a shorter and heavier head. Its skull and dentition were described as being intermediate between Canis lupus occidentalis and the dire wolf. Its skull differed from occidentalis due to its wider skull, especially at the palate, and the size of its P4 and M1 were much larger than any known timber wolf, with the P4 approaching that of the dire wolf in size.[52] It is regarded by Nowak as a taxonomic synonym of Canis lupus furlongi.[24]
Aenocyon milleri (Merriam 1918)[53] was described as a wolf different from the dire wolf by its smaller size, low sagittal crest, and a less prominent inion, but closer to the dire wolf than the timber wolf. Only one specimen was found. It is regarded by Nowak as a taxonomic synonym of Canis lupus furlongi.[24]
Canis chihliensis
| Wolf evolution – alternate proposal | |||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||
| Previously proposed evolution and branching from Eucyon towards the wolf (2008). [19][37]: p181 |
Wang and Tedford proposed that the genus Canis was the descendant of the coyote-like Eucyon davisi, and its remains first appeared in the Miocene (6 million YBP) in south-western USA and Mexico. By the Pliocene (5 million YBP), the larger Canis lepophagus appeared in the same region and by the Early Pleistocene (1 million YBP) Canis latrans (the coyote) was in existence. They proposed that the progression from Eucyon davisi to C lepophagus to the coyote was linear evolution.[18] Additionally, C. edwardii, C. latrans and C. aureus form together a small clade and because C. edwardii appeared earliest spanning the mid-Blancan (late Pliocene) to the close of the Irvingtonian (late Pleistocene) it is proposed as the ancestor.[37]: p175, 180
Nowak and Tedford also believed that it was possible for C. lupus to have been derived from a Miocene or Pliocene canid line that preceded and was separate from C. lepophagus.
In Eurasia during the Middle Pleistocene, C. falconeri gave rise to the hypercarnivore genus
Canis antonii
The large wolf C. antonii from late Pliocene to early Pleistocene China was assessed as being a variation within C. chihliensis,[37]: p197 and the large wolf C. falconeri occurred abruptly in Europe in the Early Pleistocene, perhaps representing a westward extension of C. antonii.[37]: p181
Canis borjgali
In 2020, researchers named a new species C. borjgali that was found in
Canis mosbachensis

Canis lupus
The earliest Canis lupus specimen was a fossil tooth discovered at
In France, the subspecies C. l. lunellensis Bonifay, 1971[63] discovered at Lunel-Viel, Hérault dated 400–350,000 YBP, C. l. santenaisiensis Argant, 1991[64] from Santenay, Côte-d'Or dated to 200,000 YBP, and C. lupus maximus Boudadi-Maligne, 2012[65] from Jaurens cave, Nespouls, Corrèze dated 31,000 YBP, show a progressive increase in size and are proposed to be chrono-subspecies.[66][13] In Italy, the earliest Canis lupus specimens were found at La Polledrara di Cecanibbio, 20 km north-west of Rome in strata dated 340,000–320,000 YBP.[67][68] In 2017, a study found that the dimensions of the upper and lower carnassial teeth of the early Holocene Italian wolf are close to those of C. l. maximus. Fluctuations in the size of C. lupus carnassial teeth correlate with the spread of megafauna. The Italian wolf underwent a reduction in body size with the loss of the red deer in Italy during the Renaissance.[13] The proposed lineage is:
C. etruscus → C. mosbachensis → C. l. lunellensis → C. l. santenaisiensis → C. l. maximus → C. l. lupus[13]
In 2020, a new species of wolf C. borjgali was discovered in Georgia. This wolf is proposed to be the next evolutionary step after C. mosbachensis and an ancestor of the C. lupus lineage.[69]
Canis lupus bohemica
In 2022, a new species Canis lupus bohemica was
In Hungary in 1969, a tooth (the premolar of the Maxilla) was found which dated to the Middle Pleistocene, and was assessed as being midway between that of Canis mosbachensis and the cave wolf Canis lupus spelaeus, but leaning towards C.l. spelaeus. [71] During the late Middle Pleistocene around 600,000 years ago, the Bohemian wolf diversified into two wolf lineages that specialized for different environmental and climatic conditions. One was a southern interglacial (warm climate) grey wolf of Europe which was to become the Mosbach wolf, and the other a northern glacial white wolf of Eurasia which was to become C. l. spelaeus.[70]
Canis c.f. familiaris - Paleolithic dog
There are a number of recently discovered specimens which are proposed as being Paleolithic dogs, however their taxonomy is debated. These have been found in either Europe or Siberia and date 40,000-17,000 YBP. They include
Canis familiaris (domestic dog)

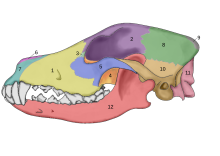
In 2002, a study was undertaken into the fossil skulls of two large canids that had been found buried within meters of the doorway of what was once a mammoth-bone hut at the Eliseevichi-I Upper Paleolithic site in the Bryansk Region on the Russian Plain, and using an accepted morphologically based definition of domestication declared them to be "Ice Age dogs". The carbon dating gave a calendar-year age estimate that ranged between 16,945 and 13,905 YBP.[74] In 2013, a study looked at one of these skulls and its mitochondrial DNA sequence was identified as Canis familiaris.[75]
In 2015, a zooarchaeologist stated that "In terms of phenotypes, dogs and wolves are fundamentally different animals."[76]
In 1986, a study of skull morphology found that the domestic dog is morphologically distinct from all other canids except the wolf-like canids. "The difference in size and proportion between some breeds are as great as those between any wild genera, but all dogs are clearly members of the same species."
Comparison with modern wolves
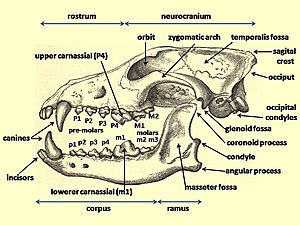
The domestic dog compared to the modern wolf shows the greatest variation in the size and shape of the skull (Evans 1979) that range from 7 to 28 cm in length (McGreevy 2004). Wolves are
Nowak indicated that orbital angle of the eye socket is an important characteristic defining the difference between the dog and the wolf, with the wolf having the lower angle. Nowak compared the orbital angles of four North American canines (including the Indian dog) and produced the following values in degrees: coyote-42.8, wolf-42.8, dog-52.9 dire wolf-53.1. The orbital angle of the eye socket was clearly larger in the dog than in the coyote and the wolf; why it was almost the same as that of the dire wolf was not commented on.[24]
Many authors have concluded that compared to the adult extant wolf, the adult domestic dog has a relatively reduced rostrum (front part of the skull), an elevated
Compared to the wolf, dog dentition is relatively less robust (Olsen 1985; Hemmer 1990), which is proposed to be due to the relaxation of natural selection when wolves became commensal scavengers, or to artificial selection (Olsen 1985; Clutton-Brock 1995). However, Kieser and Groeneveld (1992) compared the mandibulo-dental measurements of jackals (C. adustus, C. mesomelas) and Cape foxes (Vulpes chama) to equivalent-sized dogs and found that the canines of these other canids tended to be slightly smaller and their second molars larger compared to dogs, otherwise the proportions were essentially the same in all species. They concluded that "...the teeth of canids appear to have evolved in concert with one another and relatively independently of differences in dimorphism, size or functional demands". This calls into question the assumption that dog teeth are relatively small due to recent selection, suggesting that dog dentition is plesiomorphic from an ancestor that was smaller than the wolf.[80]
The reduced body size of the early dog compared to a wolf is thought due to niche selection (Olsen 1985; Morey 1992; Coppinger & Coppinger 2001). Morey (1992:199) states that "Results...are consistent with a hypothesis that early domestic dogs are evolutionary paedomorphs, products of strong selection for ontogenetically channeled size reduction and alterations of reproductive timing associated with the new domestic way of life."[80] However, in an domestication experiment the domesticated foxes remained the same size as unselected foxes (Trutt 1999:167).[76]
Wayne (1986) concluded that the dog is closer in skull morphology to C. latrans, C. aureus, C. adustus, C. mesomelas, Cuon alpinus and Lycaon pictus than to the wolf. Dahr (1942) concluded that the shape of the dog brain case is closer to that of the coyote than to that of the wolf. Manwell and Baker (1983) reviewed Dahr's work with the addition of dental data for canids and concluded that the dog ancestor was probably within the range of 13.6–20.5 kg, which is smaller than the range 27–54 kg for extant wolves (Mech 1970) and is comparable with the Dingo.[80]
The auditory bulla of the dog is relatively smaller and flatter than that of the wolf (Harrison 1973; Clutton-Brock, Corbet & Hill 1976; Nowak 1979; Olsen 1985; Wayne 1986), which is proposed to be due to relaxed selection under domestication as the dog no longer required the acute hearing of the wolf. However, bulla shape has been shown to facilitate increased sensitivity to specific frequencies but shape and size may not be correlated with acuity (Ewer 1973). Therefore, the observed difference could be that the dog bulla has retained its ancestral shape.[80]
The ventral edge of the dog's horizontal
A description of the superficial brain morphology of jackals (C. mesomelas, C. aureus), coyotes (C. latrans), wolves (C. lupus, C. rufus), and dogs indicated that the cerebellum of the dog closely approximates that of the coyote, which is closely aligned with the jackals, and that the wolves show numerous brain traits distinct from the other species (Atkins and Dillon 1971). Wolves also have serological and biochemical traits distinct from dogs (Leone and Wiens 1956; Lauer, Kuyt & Baker 1969).[80]
The tails of domestic dogs tend to curl upwards, which is not found in other canid.
During the Last Glacial Maximum, there was greater wolf genetic diversity than there is today,[9][75] and within the Pleistocene grey wolf population the variations between local environments would have encouraged a range of wolf ecotypes that were genetically, morphologically and ecologically distinct from one another.[89] One author has proposed that the most likely explanation for the different morphological characteristics of the dog compared to the wolf is that the dog's ancestor was adapted to a different niche than the wolf.[80]
Genetic record
DNA sequences
The
Mitochondrial
-
The 33,500 year old skull of the "Altai dog"
-
Location ofchromosomes of a cell nucleus
-
The structure of part of a DNAdouble helix
-
DNA molecule 1 differs from DNA molecule 2 at a single base pair location, called a single-nucleotide polymorphism (a SNP mutation)
-
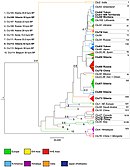
Phylogenetic tree of wolves under study
| Phylogenetic tree | |||||||||
| |||||||||
| Phylogenetic relationship between four canids.[99][100] |
The mutations that are different in these 4 sequences have been numbered and bolded. These mutations can then be used to build a phylogenetic tree for the four canids. In this example, the dog and grey wolf differ by two substitutions (highlighted in red), and each of them differs from the coyote by four substitutions.[91]
1 2 3 4 5 67
Golden Jackal A-G-C-T-G-T-C-GA-T-TC-CA
Coyote A-G-C-T-A-T-C-GA-A-TC-GA
Wolf T-G-C-T-A-T-G-GA-T-TC-CT
Dog T-G-G-T-A-T-G-GA-T-TC-CA
The mDNA sequences of the dog and wolf differ by only 0–12 substitutions within 261 base-pairs, whereas dogs always differed from coyotes and jackals by at least 20 substitutions.[91][101] This finding implies that the dog derived from the wolf and that there has been repeated back-crossing,[101] or that the dog may have descended from a now extinct species of canid whose closest living relative is the modern wolf.[102]
Marker issue
Different DNA studies may give conflicting results because of the specimens selected, the technology used, and the assumptions made by the researchers.
Timing issue
There are two key assumptions that are made for dating the divergence time for species: the generation time and the genetic mutation rate per generation. The time between generations for wolves is assumed to be three years based on the extant grey wolf, and two years for the dog based on the extant dog.[99] One recent major study assumed a generation time of 2 years for the dog for as far back as 10,000 years ago, and then assumed a generation time of 3 years (the same as the wolf) before that to calculate a proposed divergence time between the two.[9] In 2017, the wolf research scientist L. David Mech queried why evolutionary biologists were calculating the approximate time of the dog diverging from the wolf through using a wolf generation time of three years when published works using large data sets demonstrate a figure of 4.2–4.7 years. They were encouraged to recalculate their divergence dates accordingly.[109]
DNA studies are conducted but with "the mutation rate as the dominant source of uncertainty."
Wolf-like canids
The wolf-like canids (the canid subfamily
A DNA sequence alignment for the wolf-like canids gave a phylogenetic tree with the grey wolf and dog being the most closely related, followed by a close affiliation with the coyote, golden jackal and Ethiopian wolf, and the dog can hybridize in the wild with these three species. Next closest to this group are the dhole and African wild dog that both have unique meat-slicing teeth, suggesting that this adaptation was later lost by the other members.
In 2015, a study of mitochondrial genome sequences and nuclear genome sequences of African and Eurasian canids indicated that extant wolf-like canids had colonized Africa from Eurasia at least 5 times throughout the Pliocene and Pleistocene, which is consistent with fossil evidence suggesting that much of the African canid diversity resulted from the immigration of Eurasian ancestors, likely coincident with Plio-Pleistocene climatic oscillations between arid and humid conditions.[100]
The phylogenetic tree for the wolf-like canids may give conflicting positions for the black-backed jackal and the side-striped jackal relative to the genus Canis members depending on whether the genetic markers were based on mitochondrial DNA or nuclear DNA. The explanation proposed for this mito-nuclear discord is that mitochondrial DNA introgression occurred from an ancient ancestor of genus Canis into the lineage that led to the black-backed jackal around 6.2–5.2 million years ago.[120]
Phylogenetic tree of the extant wolf-like canids,[a] with the pink shading representing the species Canis lupus.
| |||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
3.5 mya |
Admixture with an extinct unknown canid
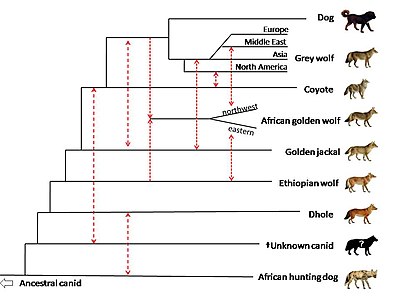
In 2018,
A genomic study on the wolves of China included museum specimens of wolves from southern China that were collected between 1963 and 1988. The wolves in the study formed 3 clades: north Asian wolves that included those from northern China and eastern Russia, Himalayan wolves from the Tibetan Plateau, and a unique population from southern China. One specimen located as far southeast as Jiangxi province shows evidence of being admixed between Tibetan-related wolves and other wolves in China. One specimen from Zhejiang province in eastern China shared gene flow with the wolves from southern China, however its genome was 12-14 percent admixed with a canid that may be the dhole or an unknown canid that predates the genetic divergence of the dhole. The wolf population from southern China is believed to be still existing in that region.[134]
Out of Beringia
Grey wolves suffered a species-wide population bottleneck (reduction) approximately 25,000 YBP during the Last Glacial Maximum. This was followed by a single population of modern wolves expanding out of a Beringia refuge to repopulate the wolf's former range, replacing the remaining Late Pleistocene wolf populations across Eurasia and North America as they did so.[12][132][133] This source population probably did not give rise to dogs, but admixed with dogs which allowed them to gain coat colour genes that are also related to immunity, and provided dogs with genes which allowed them to adapt to high-altitude environments (e.g. Tibet). This suggests that the genetic divergence of European and East Asian dogs could be based on admixture with different sub-populations of wolves.[133]
There is little genetic information available on the ancient wolves that existed prior to the bottleneck. However, studies show that one or more of these ancient populations is more directly ancestral to dogs than are modern wolves, and conceivably these were more prone to domestication by the first humans to invade Eurasia.[133]
As of 2020, the oldest known intact wolf remains belongs to a mummified pup dated 56,000 YBP that was recovered from the permafrost along a small tributary of Last Chance Creek near Dawson City, Yukon, Canada. A DNA analysis showed that it belonged to the Beringian wolf clade, that the most recent common ancestor of this clade dates to 86,700–67,500 YBP, and that this clade was basal to all other wolves except for the Himalayan wolf.[135]
Into America and Japan

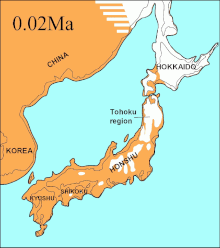
In 2016, a study built on the work of another major study
The phylogenetic tree showed the
As wolves had been in the fossil record of North America but modern wolves could trace their ancestry back only 80,000 years, the wolf haplotypes that were already in North America were replaced by these invaders, either through competitive displacement or through admixture.[131] The replacement in North America of a basal population of wolves by a more recent one supported the findings of earlier studies.[128][136][127][131] There possibly existed a panmictic wolf population with gene flow spanning Eurasia and North America until the closing of the ice sheets.[128][137][131] Once the sheets closed, the southern wolves were isolated and north of the sheets only the Beringian wolf existed. The land bridge became inundated by the sea 10,000 YBP, the sheets receded 12,000–6,000 YBP, the Beringian wolf went extinct and the southern wolves expanded to recolonize the rest of North America. All North American wolves are descended from those that were once isolated south of the ice sheets. However, much of their diversity was later lost during the twentieth century.[131]
Studies using
During the same period, the
The dog was a very successful invader of North America and had established a widespread
The wolf was exterminated in the southern part of their historic geographical range in North America by the middle of the 20th century. An mDNA study of 34 wolf remains from North America dated between 1856 and 1915 found their genetic diversity to be twice that of modern wolves in these regions, and two thirds of the haplotypes identified were unique. These results indicate that a historic population of several hundred thousand wolves once existed in Mexico and the western US.[152][130]
Divergence with the coyote
In 1993, a study proposed that the wolves of North America display skull traits more similar towards the coyote than those wolves from Eurasia.[44] In 2016, a whole-genome DNA study proposed, based on the assumptions made, that all of the North American wolves and coyotes diverged from a common ancestor less than 6,000–117,000 years ago. The study also indicated that all North American wolves have a significant amount of coyote ancestry and all coyotes some degree of wolf ancestry, and that the red wolf and eastern wolf are highly admixed with different proportions of grey wolf and coyote ancestry. However, the test that was used cannot differentiate between ancient and recent hybridization. One test indicated a wolf/coyote divergence time of 51,000 years before present that matched other studies indicating that the extant wolf came into being around this time. Another test indicated that the red wolf diverged from the coyote between 55,000 and 117,000 years before present and the Great Lakes region wolf 32,000 years before present. But a 2000 study on the haplotypes of North American canids showed that the divergences between coyotes, grey wolves, red wolves, and Algonquin (eastern) wolves happened much earlier. This study showed that there was a 3.2 % divergence between the eastern Canadian wolf and coyote gene sequences, and a 2.3 % sequence divergence between the red wolf and coyote haplotypes. However, the divergence was a much greater 8.0% between gray wolf (C. lupus) mtDNA and eastern/red wolf haplotypes, and 10.0% between gray wolf and coyote haplotypes. The sequence difference observed between eastern Canadian wolf sequences and coyote sequences is consistent with a separation of 150 000 – 300 000 years, using a divergence rate of 1–2% per 100 000 years.[153] Additionally, conflicting with the recent divergent hypothesis, the fossil record that indicates a coyote-like specimen dated to 1 million years before present.[32]
The modern grey wolf expanded out of Beringia 25,000 years ago.[133] The modern coyote appeared around 10,000 years ago, and most likely descended from the Pleistocene coyote. The most genetically basal coyote mDNA clade pre-dates the Last Glacial Maximum and is a haplotype that can only be found in the Eastern wolf. This implies that the large, wolf-like Pleistocene coyote was the ancestor of the Eastern wolf and Red wolf, which diverged from each other later in time.
Domestic dog
The domestic dog (Canis lupus familiaris) is the most widely abundant large carnivore.
In 2016, a study investigated for the first time the population subdivisions, demography, and the relationships of grey wolves based on their whole-genome sequences. The study indicated that the dog was a divergent subspecies of the grey wolf and was derived from a now-extinct ghost population of Late Pleistocene wolves,[75][9][11] and the dog and the dingo are not separate species.[11] The genome-wide phylogenetic tree indicated a genetic divergence between New World and Old World wolves, which was then followed by a divergence between the dog and Old World wolves 27,000YBP[10][11] – 29,000 YBP.[11] The dog forms a sister taxon with Eurasian grey wolves but not North American wolves. The dog had considerable pre-ancestry after its divergence from the Old World wolves before it separated into distinct lineages that are nearly as distinct from one another as they are from wolves.[11] The study suggested that previous datings based on the divergence between wolves and coyotes of one million years ago using fossils of what appeared to be coyote-like specimens may not reflect the ancestry of the modern forms.[100][9][10][11]
| Grey wolf divergence and timing in years before present |
| Whole-genome phylogenetic tree of modern grey wolf populations.[11]
|
The study indicated that the Mexican wolf was also a divergent form of grey wolf, suggesting that may have been part of an early invasion into North America.[11][152] The Tibetan wolf was found to be the most highly divergent of the Old World wolves, had suffered a historical population bottleneck and had only recently recolonized the Tibetan Plateau. Glaciation may have caused its habitat loss, genetic isolation then local adaption.[11]
The study indicated that there has been extensive
The data indicated that all wolves shared similar population trajectories, followed by population decline that coincided with the expansion of modern humans worldwide and their technology for capturing large game.[11][157] Late Pleistocene carnivores would have been social living in large prides, clans and packs in order to hunt the larger game available at that time, and these larger groups would have been more conspicuous targets for human persecutors.[157] Large dogs accompanying the humans may have accelerated the rate of decline of carnivores that competed for game,[11][158] therefore humans expanded across Eurasia, encountered wolves, domesticated some and possibly caused the decline of others.[11]
The study concluded that admixture had confounded the ability to make inferences about the place of dog domestication. Past studies based on SNPs, genome-wide similarities with Chinese wolves, and lower linkage disequilibrium might reflect regional admixture between dogs with wolves and gene flow between dog populations, with divergent dog breeds possibly maintaining more wolf ancestry in their genome. The study proposed that analysis of ancient DNA might be a better approach.[11]
In the same year, a study found that there were only 11 fixed genes that showed variation between wolves and dogs. These genes are thought to affect tameness and emotional processing ability.[159] Another study provided a listing of all of the grey wolf and dog mDNA haplotypes combined in the one phylogenetic tree.[160]
In 2018, a study compared the sequences of 61,000
Dingo
The dingo (Canis familiaris dingo) refers to the dog found in
Canis variabilis
In 2015, a study looked at the
The
The authors concluded that the structure of the modern dog gene pool was contributed to from ancient Siberian wolves and possibly from Canis c.f. variabilis.[165][166]
Rise to dominant predator
In 2015, a study looked at the paleoecology of large carnivores across the
Wolf population differences
The grey wolf Canis lupus is a highly adaptable species that is able to exist in a range of environments and which possesses a wide distribution across the
Apart from domestication, humans have harmed the wolf by restricting its habitat through persecution. This has caused a dramatic decrease in its population size over the last two centuries.[171][172] The shrinking of its habitats that overlap with those of close-relatives such as dogs and coyotes have led to numerous occurrences of hybridization.[173][174] These events, in addition to recent turnovers (extinctions and repopulations by other geneotypes), has made the unravelling of the phylogeographic history of the wolf difficult.[122]
Ecotypes
An ecotype is a variant in which the phenotypic differences are too few or too subtle to warrant being classified as a subspecies. These can occur in the same geographic region where distinct habitats such as meadow, forest, swamp, and sand dunes provide ecological niches. Where similar ecological conditions occur in widely separated places it is possible for a similar ecotype to occur. This is different from a subspecies, which may exist across a number of different habitats. In animals, ecotypes can be regarded as micro-subspecies that owe their differing characteristics to the effects of a very local environment.[175] Ecotypes have no taxonomic rank.
Grey wolves have a wide, natural distribution across the
In 2013, a genetic study found that the wolf population in Europe was divided along a north–south axis and formed five major clusters. Three clusters were identified occupying southern and central Europe in Italy, the Carpathians, and the Dinaric-Balkans. Another two clusters were identified occupying north-central Europe and the Ukrainian steppe. The Italian wolf consisted of an isolated population with low genetic diversity. Wolves from Croatia, Bulgaria, and Greece formed the Dinaric-Balkans cluster. Wolves from Finland, Latvia, Belarus, Poland and Russia formed the north-central Europe cluster with wolves from the Carpathians cluster a mixture of wolves from the north-central cluster and the Dinaric-Balkans cluster. The wolves from the Carpathians were more similar to the wolves from the Ukrainian Steppe than they were to wolves from north-central Europe. These clusters may have been the result of expansion from glacial refugia, an adaptation to local environments, and landscape fragmentation and the killing of wolves in some areas by humans.[180]
In 2016, two studies compared the sequences of 42,000
Ecological factors including habitat type, climate, prey specialization and predatory competition will greatly influence grey wolf genetic population structure and cranio-dental plasticity.[184][89][7][2][185][186][4][5][6] During the Last Glacial Maximum, there was greater wolf genetic diversity than there is today,[9][75] and within the Pleistocene grey wolf population the variations between local environments would have encouraged a range of wolf ecotypes that were genetically, morphologically and ecologically distinct from one another.[89]
Pleistocene wolves
During the
There are a small number of Canis remains that have been found at Goyet Cave, Belgium (36,500 YBP)[73] Razboinichya Cave, Russia (33,500 YBP)[190] Kostenki 8, Russia (33,500–26,500 YBP)[191] Predmosti, Czech Republic (31,000 YBP)[192] and Eliseevichi 1, Russia (17,000 YBP).[74] Based on cranial morphometric study of the characteristics thought to be associated with the domestication process, these have been proposed as early Paleolithic dogs.[191] These characteristics of shortened rostrum, tooth crowding, and absence or rotation of premolars have been documented in both ancient and modern wolves.[127][89][6][193][194][195] Rather than representing early dogs, these specimens may represent "a morphologically distinct local, now extinct, population of wolves".[89][196]
Notes
- ^ Phylogenetic relationships and time of genetic divergence for the extant wolf-like canids based on DNA of the entire cell nucleus.[121] Exception 1: Side-striped jackal and Black-backed jackal added based on segments of DNA taken from the cell nucleus[99][100] with proposed divergence times in millions of years.[100] Exception 2: The position of the Himalayan wolf is based on segments of mitochondrial DNA[100][122][123][124] and segments from the cell nucleus.[100][123][124] Exception 3: The position of the Indian plains wolf is based on mDNA.[125][126][127][128][129][130] and its timing 120,000 years ago.[12] Exception 4: Late Pleistocene wolf included for time-line purposes only - the most recent common ancestor for all other Canis lupus specimens – modern and extinct but excluding the Indian plains wolf and the Himalayan wolf – was 80,000 years before present.[75][131] Exception 5: Grey wolf and dog timing 25,000 years ago.[12][132][133]
References
- ^ Dawkins, William Boyd; Sanford, W. Ayshford; Reynolds, Sidney H. (1912). "British Pleistocene Hyænidæ, Ursidæ, Canidæ, and Mustelidæ". A Monograph of the British Pleistocene Mammalia. Vol. 2. London: Palaeontographical Society.
- ^ S2CID 14459019.
- ^ S2CID 7808556.
- ^ S2CID 4840903.
- ^ S2CID 11864260.
- ^ .
- ^ a b c d e f g h i j k l m n o p Leonard, Jennifer (2014). "Ecology drives evolution in grey wolves" (PDF). Evolution Ecology Research. 16: 461–473.
- ^ .
- ^ PMID 24453982.
- ^ PMID 26004765.
- ^ PMID 26680994.
- ^ PMID 31840921.
- ^ .
- ^ Wang & Tedford 2008, p. 8.
- ^ Wang & Tedford 2008, p. 1.
- ^ Wang & Tedford 2008, p. 116.
- ^ "Eucyon davisi". Fossilworks. Retrieved 17 December 2021.
- ^ a b Wang & Tedford 2008, p. 58.
- ^ a b c d Wang & Tedford 2008, p. 148.
- ^ Vislobokova, I., Sotnikova, M. & Dodonov, A., 2003 – Bio-events and diversity of the Late Miocene-Pliocene mammal faunas of Russia and adjacent areas – in: Reumer, J.W.F. & Wessels, W. (eds.) – DISTRIBUTION AND MIGRATION OF TERTIARY MAMMALS IN EURASIA. A VOLUME IN HONOUR OF HANS DE BRUIJN – DEINSEA 10: 563–574 [ISSN 0923-9308] Published 1 December 2003
- .
- .
- ^ ISBN 978-0-226-51696-7.
- ^ ISBN 978-0-89338-007-6.
- ^ Wayne, R. K. (1995). "Red wolf: To conserve or not to conserve" (PDF). In Macdonald, D.; Handoca, L. (eds.). Canid News. Vol. 3. Newsletter of the IUCN/SSC Specialist Group. pp. 7–12.
- ^ Hoffmeister, D. F., and W. W. Goodpaster. 1954. The mammals of the Huachuca Mountains, southeastern Arizona. Illinois Biological Monographs 24:1— 152.
- JSTOR 3881428.
- ISBN 978-0-442-22430-1.
- ^ ISBN 978-0-8130-0361-0.
- ISBN 978-0-8130-0361-0.
- S2CID 4364642.
- ^ a b Wang, Xiaoming; Tedford, Richard H.; Dogs: Their Fossil Relatives and Evolutionary History. New York: Columbia University Press, 2008.
- ^ "Canis lepophagus". Fossilworks. Retrieved 17 December 2021.
- ^ Kurten, B. (1974). "A History of Coyote-Like Dogs (Canidae, Mamalia)". Acta Zoologica Fennica (140): 1–38.
- ^ ISBN 978-0-231-03733-4.
- ^ .
- ^ S2CID 83594819.
- ^ Berta, A. (1995). "Fossil carnivores from the Leisey Shell Pits, Hillsborough County, Florida". In Hulbert, R. C. Jr.; Morgan, G. S.; Webb, S. D. (eds.). Paleontology and geology of the Leisey Shell Pits, early Pleistocene of Florida. Vol. 37. Bulletin of the Florida Museum of Natural History. pp. 463–499.
- ^ "Canis ambrusteri". Fossilworks. Retrieved 17 December 2021.
- ^ "Canis dirus". Fossilworks. Retrieved 17 December 2021.
- ^ Kurten, B. 1984: Geographic differentiation in the Rancholabrean dire wolf (Canis dirus Leidy) in North America. In Genoways, H. H. & Dawson, M. R. (eds.): Contributions in Quaternary Vertebrate Paleontology: A Volume in Memorial to John E. Guilday, 218–227. Carnegie Museum of Natural History Special Publication 8
- ISBN 978-0-8117-0496-0.
- doi:10.1139/z87-171.
- ^ a b Goulet, G. D. (1993). Comparison of temporal and geographical skull variation among Nearctic, modern, Holocene, and late Pleistocene gray wolves (Canis lupus) and selected Canis (Master's Degree). Winnipeg: University of Manitoba. pp. 1–116.
- ISBN 978-0-520-09960-9.
- ^ Wang & Tedford 2008, p. 52.
- S2CID 366077.
- S2CID 231604957.
- ^ a b Canis occidentalis furlongi Merriam (1910), Univ. California Publ. Bull. Dept. Geol., 5:393
- ISBN 978-0-12-319250-9.
- ^ Canis milleri Merriam (1912) Mem. Univ. California, 1:247
- ^ Merriam, J. C. (1912). "The fauna of Rancho La Brea, Part II. Canidae". Memoirs of the University of California. 1: 217–273.
- ^ Aenocyon milleri Merriam (1918) Univ. California Publ. Bull. Dept. Geol., 10:533.
- ^ Tedford, R.H. & Qiu, Z.-X., 1996 - A new canid genus from the Pliocene of Yushe, Shanxi Province – Vertebrata PalAsiatica 34 (1): 27–40
- ^ Tong; Hu, N.; Wang, X. (2012). "New remains of Canis chihliensis (Mammalia, Carnivora) from Shanshenmiaozui, a lower Pleistocene site in Yangyuan, Hebei". Vertebrata PalAsiatica. 50 (4): 335–360. Archived from the original on 2016-09-19. Retrieved 2016-09-19.
- ^ Wang & Tedford 2008, p. 61.
- ^ Wang & Tedford 2008, p. 105.
- ^ Wang & Tedford 2008, p. 149.
- ^ Wang & Tedford 2008, p. 60.
- ^ ISSN 2296-6463.
- ^ ISBN 978-0-202-30953-8.
- S2CID 140572760.
- ^ Bonifay, M.F., 1971. Carnivores quaternaires du Sud-Est de la France. Muséum national d'Histoire naturelle, Paris, 334 p. (Mémoires du Muséum national d'Histoire naturelle, Sér. C – Sciences de la Terre (1950–1992) ; 21 (2)). [Quaternary carnivores of the South-East of France]
- ^ Argant A., 1991 – Carnivores quaternaires de Bourgogne. Documents du Laboratoire de Géologie de Lyon, n° 115, 301 p.
- .
- .
- .
- .
- S2CID 216337804.
- ^ .
- ISBN 978-0444556356.
- ISBN 978-3-030-04752-8.
- ^ .
- ^ a b Sablin, Mikhail V.; Khlopachev, Gennady A. (2002). "The Earliest Ice Age Dogs: Evidence from Eliseevichi I" (PDF). Zoological Institute of the Russian Academy of Sciences. Wenner-Gren Foundation for Anthropological Research. Retrieved 10 January 2015.
- ^ S2CID 1526260.
- ^ .
- PMID 28556057.
- S2CID 26967649.
- PMID 20668685.
- ^ S2CID 14850835. Archived from the original(PDF) on 2019-09-20. Retrieved 2019-09-20.
- ^ Drake, Abby Grace, "Evolution and development of the skull morphology of canids: An investigation of morphological integration and heterochrony" (January 1, 2004). Doctoral Dissertations Available from Proquest. Paper AAI3136721. link
- S2CID 20893501.
- ISBN 978-0-521-41529-3.
- ^ Wang & Tedford 2008, p. 158.
- ISBN 978-0-521-41529-3.
- JSTOR 3503924. Archived from the original(PDF) on 24 September 2015. Retrieved 4 June 2015.
- PMID 26893534.
- ^ Elliot, D.G., and M. Wong. 1972. "Acid phosphatase, handy enzyme that separates the dog from the wolf". Acta Biologica et Medica Germanica 28:957–962
- ^ .
- .
- ^ ISBN 978-0-520-24638-6.
- S2CID 205016024.
- ISBN 978-0-87893-463-8.[page needed]
- S2CID 1665621.
- S2CID 11567406.
- PMID 11700280.
- ISBN 978-0-412-03781-8.
- ISBN 978-0-674-66638-2.
- ^ PMID 16341006.
- ^ PMID 26234211.
- ^ PMID 9180076.
- ^ S2CID 5547543.
- PMID 19666600.
- PMID 19723671.
- ^ PMID 25429025.
- PMID 16213754.
- PMID 20181231.
- S2CID 16003335.
- S2CID 206647686.
- S2CID 206647686.
- ^ PMID 8337763.
- ISBN 978-0-412-34360-5.
- ISBN 978-1-84593-940-3.
- PMID 7151489.
- S2CID 25976335.
- ^ Pocock, R. I. (1941). Fauna of British India: Mammals. Vol. 2. Taylor & Francis. pp. 146–163.
- S2CID 24540274.
- PMID 1968637. Retrieved 21 December 2011.
- ISBN 978-0-12407-931-1.
- PMID 33591975.
- ^ PMID 30344120.
- ^ .
- ^ PMID 28680672.
- ^ hdl:10651/50748.
- .
- PMID 15101402.
- ^ S2CID 14039133. Archived from the original(PDF) on 2016-12-28. Retrieved 2016-06-11.
- ^ PMID 20409299.
- PMID 21298107.
- ^ ISBN 978-0-19-954566-7.
- ^ S2CID 88740690.
- ^ hdl:10651/50748.
- ^ PMID 32286714.
- PMID 31563851.
- PMID 33352124.
- .
- PMID 17686436.
- S2CID 12896064.
- .
- PMID 21573241.
- PMID 29713682.
- ^ PMID 25132126.
- Shōbara: Hiba Society of Natural History. pp. 89–104.
- S2CID 11569628.
- ^ Walker, Brett (2008). The Lost Wolves of Japan. University of Washington Press. p. 42.
- ^ S2CID 27005517.
- PMID 33364590.
- ^ .
- PMID 25532803.
- ^ Wolf and Man. Evolution in Parallel. R.L. Hall and H.S. Sharp. Academic Press, New York, 1978
- ^ Olsen, S. J. (1985). Origins of the domestic dog: the fossil record. Univ. of Arizona Press, Tucson, USA. pp. 88–89.
- ^ S2CID 11343074.
- ^ Wilson, Paul; Grewal, Sonya; Lawford, Ian; Heal, Jennifer (December 2000). "DNA profiles of the eastern Canadian wolf and the red wolf provide evidence for a common evolutionary history of the gray wolf". ResearchGate. Canadian Journal of Zoology. Retrieved November 28, 2022.
- S2CID 31680635.
- ^ PMID 24453989.
- S2CID 90588969.
- ^ PMID 26504224.
- ISBN 978-0-674-73676-4.
- PMID 26754411.
- PMID 27042801.
- PMID 29875809.
- PMID 15299143.
- ^ PMID 21900326.
- PMID 22618967.
- ^ PMID 26018528.
- ISBN 978-3-030-04587-6.
- S2CID 129706533.
- PMID 23658198.
- ^ .
- .
- S2CID 11343074.
- PMID 21566151.
- S2CID 1161310.
- ISBN 978-0-674-86250-0.
- PMID 12590776.
- ^ S2CID 38788935.
- S2CID 85815288.
- .
- PMID 24146871.
- ^ S2CID 17798894.
- .
- S2CID 20760315.
- PMID 20409351.
- S2CID 29313917.
- ^ Carmichael, L. E. (2006). Ecological Genetics of Northern Wolves and Arctic Foxes. Ph.D. Dissertation (PDF). University of Alberta.
- .
- from the original on August 2, 2017. Retrieved December 23, 2014.
- PMID 24346500.
- PMID 21829526.
- ^ .
- .
- ^ Boudadi-Maligne, M. (2010). "Les Canis pleistocenes du sud de la France: approche biosystematique, evolutive et biochronologique. Ph.D. dissertation" (PDF). Archéologie et Préhistoire (4126). Universitie Bordeaux.
- doi:10.1002/oa.2260.
- .
- PMID 22615366.
Works
- OCLC 502410693.
















