Protein
Proteins are large
A linear chain of amino acid residues is called a
Once formed, proteins only exist for a certain period and are then degraded and recycled by the cell's machinery through the process of protein turnover. A protein's lifespan is measured in terms of its half-life and covers a wide range. They can exist for minutes or years with an average lifespan of 1–2 days in mammalian cells. Abnormal or misfolded proteins are degraded more rapidly either due to being targeted for destruction or due to being unstable.
Like other biological macromolecules such as polysaccharides and nucleic acids, proteins are essential parts of organisms and participate in virtually every process within cells. Many proteins are enzymes that catalyse biochemical reactions and are vital to metabolism. Proteins also have structural or mechanical functions, such as actin and myosin in muscle and the proteins in the cytoskeleton, which form a system of scaffolding that maintains cell shape. Other proteins are important in cell signaling, immune responses, cell adhesion, and the cell cycle. In animals, proteins are needed in the diet to provide the essential amino acids that cannot be synthesized. Digestion breaks the proteins down for metabolic use.
Proteins may be
History and etymology
Proteins were recognized as a distinct class of biological molecules in the eighteenth century by
Proteins were first described by the Dutch chemist
Early nutritional scientists such as the German
The difficulty in purifying proteins in large quantities made them very difficult for early protein biochemists to study. Hence, early studies focused on proteins that could be purified in large quantities, e.g., those of blood, egg white, various toxins, and digestive/metabolic enzymes obtained from slaughterhouses. In the 1950s, the
The first protein to be sequenced was insulin, by Frederick Sanger, in 1949. Sanger correctly determined the amino acid sequence of insulin, thus conclusively demonstrating that proteins consisted of linear polymers of amino acids rather than branched chains, colloids, or cyclols.[16] He won the Nobel Prize for this achievement in 1958.[17]

With the development of
Since then,
Number of proteins encoded in genomes
The number of proteins encoded in a genome roughly corresponds to the number of genes (although there may be a significant number of genes that encode RNA of protein, e.g. ribosomal RNAs). Viruses typically encode a few to a few hundred proteins, archaea and bacteria a few hundred to a few thousand, while eukaryotes typically encode a few thousand up to tens of thousands of proteins (see genome size for a list of examples).
Classification
Proteins are primarily classified by sequence and structure, although other classifications are commonly used. Especially for enzymes the EC number system provides a functional classification scheme. Similarly, the gene ontology classifies both genes and proteins by their biological and biochemical function, but also by their intracellular location.
Sequence similarity is used to classify proteins both in terms of evolutionary and functional similarity. This may use either whole proteins or protein domains, especially in multi-domain proteins. Protein domains allow protein classification by a combination of sequence, structure and function, and thy can be combined in many different ways. In an early study of 170,000 proteins, about two-thirds were assigned at least one domain, with larger proteins containing more domains (e.g. proteins larger than 600 amino acids having an average of more than 5 domains).[24]
Biochemistry
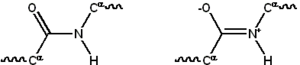
Most proteins consist of linear
The peptide bond has two
The words protein, polypeptide, and
Interactions
Proteins can interact with many types of molecules, including
Abundance in cells
It has been estimated that average-sized
Synthesis
Biosynthesis


Proteins are assembled from amino acids using information encoded in genes. Each protein has its own unique amino acid sequence that is specified by the
The process of synthesizing a protein from an mRNA template is known as
The size of a synthesized protein can be measured by the number of amino acids it contains and by its total
Chemical synthesis
Short proteins can also be synthesized chemically by a family of methods known as
Structure

Most proteins
- amino acid sequence. A protein is a polyamide.
- Secondary structure: regularly repeating local structures stabilized by hydrogen bonds. The most common examples are the α-helix, β-sheet and turns. Because secondary structures are local, many regions of different secondary structure can be present in the same protein molecule.
- post-translational modifications. The term "tertiary structure" is often used as synonymous with the term fold. The tertiary structure is what controls the basic function of the protein.
- Quaternary structure: the structure formed by several protein molecules (polypeptide chains), usually called protein subunits in this context, which function as a single protein complex.
- Quinary structure: the signatures of protein surface that organize the crowded cellular interior. Quinary structure is dependent on transient, yet essential, macromolecular interactions that occur inside living cells.
Proteins are not entirely rigid molecules. In addition to these levels of structure, proteins may shift between several related structures while they perform their functions. In the context of these functional rearrangements, these tertiary or quaternary structures are usually referred to as "

Proteins can be informally divided into three main classes, which correlate with typical tertiary structures:
A special case of intramolecular hydrogen bonds within proteins, poorly shielded from water attack and hence promoting their own
Protein domains
Many proteins are composed of several protein domains, i.e. segments of a protein that fold into distinct structural units. Domains usually also have specific functions, such as enzymatic activities (e.g. kinase) or they serve as binding modules (e.g. the SH3 domain binds to proline-rich sequences in other proteins).
Sequence motif
Short amino acid sequences within proteins often act as recognition sites for other proteins.[42] For instance, SH3 domains typically bind to short PxxP motifs (i.e. 2 prolines [P], separated by two unspecified amino acids [x], although the surrounding amino acids may determine the exact binding specificity). Many such motifs has been collected in the Eukaryotic Linear Motif (ELM) database.
Protein topology
Topology of a protein describes the entanglement of the backbone and the arrangement of contacts within the folded chain.[43] Two theoretical frameworks of knot theory and Circuit topology have been applied to characterise protein topology. Being able to describe protein topology opens up new pathways for protein engineering and pharmaceutical development, and adds to our understanding of protein misfolding diseases such as neuromuscular disorders and cancer.
Cellular functions
Proteins are the chief actors within the cell, said to be carrying out the duties specified by the information encoded in genes.[28] With the exception of certain types of RNA, most other biological molecules are relatively inert elements upon which proteins act. Proteins make up half the dry weight of an Escherichia coli cell, whereas other macromolecules such as DNA and RNA make up only 3% and 20%, respectively.[44] The set of proteins expressed in a particular cell or cell type is known as its proteome.

The chief characteristic of proteins that also allows their diverse set of functions is their ability to bind other molecules specifically and tightly. The region of the protein responsible for binding another molecule is known as the
Proteins can bind to other proteins as well as to small-molecule substrates. When proteins bind specifically to other copies of the same molecule, they can oligomerize to form fibrils; this process occurs often in structural proteins that consist of globular monomers that self-associate to form rigid fibers. Protein–protein interactions also regulate enzymatic activity, control progression through the cell cycle, and allow the assembly of large protein complexes that carry out many closely related reactions with a common biological function. Proteins can also bind to, or even be integrated into, cell membranes. The ability of binding partners to induce conformational changes in proteins allows the construction of enormously complex signaling networks.[31]: 830–49 As interactions between proteins are reversible, and depend heavily on the availability of different groups of partner proteins to form aggregates that are capable to carry out discrete sets of function, study of the interactions between specific proteins is a key to understand important aspects of cellular function, and ultimately the properties that distinguish particular cell types.[46][47]
Enzymes
The best-known role of proteins in the cell is as
The molecules bound and acted upon by enzymes are called
Dirigent proteins are members of a class of proteins that dictate the stereochemistry of a compound synthesized by other enzymes.[51]
Cell signaling and ligand binding
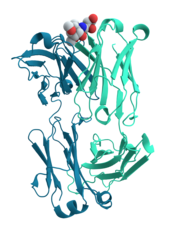
Many proteins are involved in the process of
Many ligand transport proteins bind particular
Structural proteins
Structural proteins confer stiffness and rigidity to otherwise-fluid biological components. Most structural proteins are
Other proteins that serve structural functions are motor proteins such as myosin, kinesin, and dynein, which are capable of generating mechanical forces. These proteins are crucial for cellular motility of single celled organisms and the sperm of many multicellular organisms which reproduce sexually. They also generate the forces exerted by contracting muscles[31]: 258–64, 272 and play essential roles in intracellular transport.
Protein evolution
A key question in molecular biology is how proteins evolve, i.e. how can
Methods of study
The activities and structures of proteins may be examined in vitro, in vivo, and in silico. In vitro studies of purified proteins in controlled environments are useful for learning how a protein carries out its function: for example, enzyme kinetics studies explore the chemical mechanism of an enzyme's catalytic activity and its relative affinity for various possible substrate molecules. By contrast, in vivo experiments can provide information about the physiological role of a protein in the context of a cell or even a whole organism. In silico studies use computational methods to study proteins.
Protein purification
To perform
For natural proteins, a series of purification steps may be necessary to obtain protein sufficiently pure for laboratory applications. To simplify this process, genetic engineering is often used to add chemical features to proteins that make them easier to purify without affecting their structure or activity. Here, a "tag" consisting of a specific amino acid sequence, often a series of histidine residues (a "His-tag"), is attached to one terminus of the protein. As a result, when the lysate is passed over a chromatography column containing nickel, the histidine residues ligate the nickel and attach to the column while the untagged components of the lysate pass unimpeded. A number of different tags have been developed to help researchers purify specific proteins from complex mixtures.[58]
Cellular localization
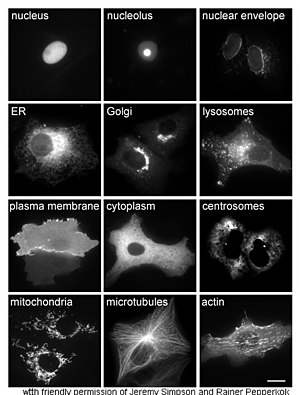
The study of proteins in vivo is often concerned with the synthesis and localization of the protein within the cell. Although many intracellular proteins are synthesized in the
Other methods for elucidating the cellular location of proteins requires the use of known compartmental markers for regions such as the ER, the Golgi, lysosomes or vacuoles, mitochondria, chloroplasts, plasma membrane, etc. With the use of fluorescently tagged versions of these markers or of antibodies to known markers, it becomes much simpler to identify the localization of a protein of interest. For example,
Other possibilities exist, as well. For example,
Finally, the gold-standard method of cellular localization is
Through another genetic engineering application known as site-directed mutagenesis, researchers can alter the protein sequence and hence its structure, cellular localization, and susceptibility to regulation. This technique even allows the incorporation of unnatural amino acids into proteins, using modified tRNAs,[64] and may allow the rational design of new proteins with novel properties.[65]
Proteomics
The total complement of proteins present at a time in a cell or cell type is known as its proteome, and the study of such large-scale data sets defines the field of proteomics, named by analogy to the related field of genomics. Key experimental techniques in proteomics include 2D electrophoresis,[66] which allows the separation of many proteins, mass spectrometry,[67] which allows rapid high-throughput identification of proteins and sequencing of peptides (most often after in-gel digestion), protein microarrays, which allow the detection of the relative levels of the various proteins present in a cell, and two-hybrid screening, which allows the systematic exploration of protein–protein interactions.[68] The total complement of biologically possible such interactions is known as the interactome.[69] A systematic attempt to determine the structures of proteins representing every possible fold is known as structural genomics.[70]
Structure determination
Discovering the tertiary structure of a protein, or the quaternary structure of its complexes, can provide important clues about how the protein performs its function and how it can be affected, i.e. in
Many more gene sequences are known than protein structures. Further, the set of solved structures is biased toward proteins that can be easily subjected to the conditions required in
Structure prediction

Complementary to the field of structural genomics, protein structure prediction develops efficient mathematical models of proteins to computationally predict the molecular formations in theory, instead of detecting structures with laboratory observation.[75] The most successful type of structure prediction, known as homology modeling, relies on the existence of a "template" structure with sequence similarity to the protein being modeled; structural genomics' goal is to provide sufficient representation in solved structures to model most of those that remain.[76] Although producing accurate models remains a challenge when only distantly related template structures are available, it has been suggested that sequence alignment is the bottleneck in this process, as quite accurate models can be produced if a "perfect" sequence alignment is known.[77] Many structure prediction methods have served to inform the emerging field of protein engineering, in which novel protein folds have already been designed.[78] Also proteins (in eukaryotes ~33%) contain large unstructured but biologically functional segments and can be classified as intrinsically disordered proteins.[79] Predicting and analysing protein disorder is, therefore, an important part of protein structure characterisation.[80]
Bioinformatics
A vast array of computational methods have been developed to analyze the structure, function and evolution of proteins. The development of such tools has been driven by the large amount of genomic and proteomic data available for a variety of organisms, including the
In silico simulation of dynamical processes
A more complex computational problem is the prediction of intermolecular interactions, such as in
Beyond classical molecular dynamics,
Chemical analysis
The total nitrogen content of organic matter is mainly formed by the amino groups in proteins. The Total Kjeldahl Nitrogen (
Nutrition
Most
In animals, amino acids are obtained through the consumption of foods containing protein. Ingested proteins are then broken down into amino acids through digestion, which typically involves denaturation of the protein through exposure to acid and hydrolysis by enzymes called proteases. Some ingested amino acids are used for protein biosynthesis, while others are converted to glucose through gluconeogenesis, or fed into the citric acid cycle. This use of protein as a fuel is particularly important under starvation conditions as it allows the body's own proteins to be used to support life, particularly those found in muscle.[90]
In animals such as dogs and cats, protein maintains the health and quality of the skin by promoting hair follicle growth and keratinization, and thus reducing the likelihood of skin problems producing malodours.[91] Poor-quality proteins also have a role regarding gastrointestinal health, increasing the potential for flatulence and odorous compounds in dogs because when proteins reach the colon in an undigested state, they are fermented producing hydrogen sulfide gas, indole, and skatole.[92] Dogs and cats digest animal proteins better than those from plants, but products of low-quality animal origin are poorly digested, including skin, feathers, and connective tissue.[92]
See also
References
- archive.org
- ^ Mulder GJ (1838). "Sur la composition de quelques substances animales". Bulletin des Sciences Physiques et Naturelles en Néerlande: 104.
- S2CID 4271525.
- ^ S2CID 32843102.
- ^ New Oxford Dictionary of English
- ISBN 978-0-19-860694-9.
- ^ Reynolds and Tanford (2003).
- ^ Bischoff TL, Voit C (1860). Die Gesetze der Ernaehrung des Pflanzenfressers durch neue Untersuchungen festgestellt (in German). Leipzig, Heidelberg.
{{cite book}}: CS1 maint: location missing publisher (link) - ^ "Hofmeister, Franz". encyclopedia.com. Archived from the original on 5 April 2017. Retrieved 4 April 2017.
- ^ "Protein, section: Classification of protein". britannica.com. Archived from the original on 4 April 2017. Retrieved 4 April 2017.
- .
- PMID 14834145.
- PMID 13332017.
- PMID 14404936.
- PMID 14363272.
- PMID 15396627.
- ^ Sanger F. (1958), Nobel lecture: The chemistry of insulin (PDF), Nobelprize.org, archived (PDF) from the original on 2013-03-19, retrieved 2016-02-09
- ^ .
- S2CID 4257461.
- S2CID 4162786.
- PMID 18403197.
- PMID 18393863.
- ^ "Summary Statistics". RCSB PDB. Retrieved 2024-04-20.
- PMID 15808866.
- ^ Nelson DL, Cox MM (2005). Lehninger's Principles of Biochemistry (4th ed.). New York, New York: W. H. Freeman and Company.
- PMID 16214343.
- ^ ISBN 978-0-07-146197-9.
- ^ a b c Lodish H, Berk A, Matsudaira P, Kaiser CA, Krieger M, Scott MP, Zipurksy SL, Darnell J (2004). Molecular Cell Biology (5th ed.). New York, New York: WH Freeman and Company.
- PMID 28723063.
- ^ ISBN 978-0-8153-2305-1.
- ^ ISBN 978-0-8053-3931-4.
- PMID 24114984.
- PMID 22068332.
- PMID 23676674.
- ISBN 978-0-19-963789-8.
- PMID 27789699.
- S2CID 20237314.
- PMID 14965208.
- PMID 16226484.
- S2CID 5432012.
- PMID 12944304.
- PMID 21909575.
- S2CID 218957613.
- ^ a b Voet D, Voet JG. (2004). Biochemistry Vol 1 3rd ed. Wiley: Hoboken, NJ.
- PMID 11732604.
- S2CID 205469320.
- PMID 19273120.
- PMID 10592255.
- PMID 7809611.
- ^ EBI External Services (2010-01-20). "The Catalytic Site Atlas at The European Bioinformatics Institute". Ebi.ac.uk. Archived from the original on 2013-08-03. Retrieved 2011-01-16.
- S2CID 1896003.
- PMID 10702616.
- ISBN 978-0-470-01617-6.
- PMID 25157146.
- ^ PMID 30962359.
- PMID 28545359.
- PMID 18369866.
- S2CID 206934268.
- PMID 18691124.
- S2CID 205418407.
- PMID 10610805.
- ISBN 978-0-521-65873-7.
- PMID 18553098.
- PMID 12470735.
- PMID 10981626.
- S2CID 28594824.
- PMID 18806738.
- PMID 18218650.
- PMID 18839074.
- PMID 12547423.
- PMID 16319884.
- PMID 18430752.
- PMID 15059248.
- PMID 22130980.
- PMID 18436442.
- PMID 16787261.
- PMID 15653774.
- S2CID 1939390.
- PMID 15019783.
- ISBN 978-1-4200-7893-0.[page needed]
- PMID 18336319.
- PMID 12417204.
- S2CID 1477100.
- PMID 16910676.
- PMID 29216421.
- PMID 23105920.
- PMID 17034338.
- PMID 23959242.
- doi:10.4141/S01-054.
- PMID 12771367.
- PMID 9868266.
- ^ a b Case LP, Daristotle L, Hayek MG, Raasch MF (2010). Canine and Feline Nutrition-E-Book: A Resource for Companion Animal Professionals. Elsevier Health Sciences.
Further reading
- Textbooks
- Branden C, Tooze J (1999). Introduction to Protein Structure. New York: Garland Pub. ISBN 978-0-8153-2305-1.
- Murray RF, Harper HW, Granner DK, Mayes PA, Rodwell VW (2006). Harper's Illustrated Biochemistry. New York: Lange Medical Books/McGraw-Hill. ISBN 978-0-07-146197-9.
- Van Holde KE, Mathews CK (1996). Biochemistry. Menlo Park, California: Benjamin/Cummings Pub. Co., Inc. ISBN 978-0-8053-3931-4.
External links
Databases and projects
- NCBI Entrez Protein database
- NCBI Protein Structure database
- Human Protein Reference Database
- Human Proteinpedia
- Folding@Home (Stanford University) Archived 2012-09-08 at the Wayback Machine
- Protein Databank in Europe (see also PDBeQuips, short articles and tutorials on interesting PDB structures)
- Research Collaboratory for Structural Bioinformatics (see also Molecule of the Month Archived 2020-07-24 at the Wayback Machine, presenting short accounts on selected proteins from the PDB)
- Proteopedia – Life in 3D: rotatable, zoomable 3D model with wiki annotations for every known protein molecular structure.
- UniProt the Universal Protein Resource
Tutorials and educational websites
- "An Introduction to Proteins" from HOPES(Huntington's Disease Outreach Project for Education at Stanford)
- Proteins: Biogenesis to Degradation – The Virtual Library of Biochemistry and Cell Biology
