CCR5
Ensembl | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| UniProt | |||||||||
| RefSeq (mRNA) | |||||||||
| RefSeq (protein) | |||||||||
| Location (UCSC) | Chr 3: 46.37 – 46.38 Mb | Chr 9: 123.92 – 123.95 Mb | |||||||
| PubMed search | [3] | [4] | |||||||
| View/Edit Human | View/Edit Mouse |
C-C chemokine receptor type 5, also known as CCR5 or CD195, is a
In humans, the CCR5 gene that encodes the CCR5 protein is
Function
The CCR5 protein belongs to the
.CCR5's cognate ligands include CCL3, CCL4 (also known as MIP 1α and 1β, respectively), and CCL3L1.[14][15] CCR5 furthermore interacts with CCL5 (a chemotactic cytokine protein also known as RANTES).[14][16][17]
CCR5 is predominantly
Modulation of CCR5 activity contributes to a non-pathogenic course of infection with simian immunodeficiency virus (SIV) in several African non-human primate species that are long-term natural hosts of SIV and avoid immunodeficiency upon the infection.[23] These regulatory mechanisms include: genetic deletions that abrogate cell surface expression of CCR5,[24] downregulation of CCR5 on the surface of CD4+ T cells, in particular on memory cells,[25] and delayed onset of CCR5 expression on the CD4+ T cells during development.[26][27]

HIV
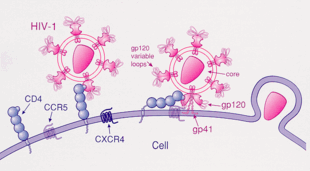
HIV-1 most commonly uses the chemokine receptors CCR5 and/or CXCR4 as co-receptors to enter target immunological cells.[28] These receptors are located on the surface of host immune cells whereby they provide a method of entry for the HIV-1 virus to infect the cell.[29] The HIV-1 envelope glycoprotein structure is essential in enabling the viral entry of HIV-1 into a target host cell.[29] The envelope glycoprotein structure consists of two protein subunits cleaved from a Gp160 protein precursor encoded for by the HIV-1 env gene: the Gp120 external subunit and the Gp41 transmembrane subunit.[29] This envelope glycoprotein structure is arranged into a spike-like structure located on the surface of the virion and consists of a trimer of Gp120-Gp41 hetero-dimers.[29] The Gp120 envelope protein is a chemokine mimic.[28] Though it lacks the unique structure of a chemokine, it is still capable of binding to the CCR5 and CXCR4 chemokine receptors.[28] During HIV-1 infection, the Gp120 envelope glycoprotein subunit binds to a CD4 glycoprotein and a HIV-1 co-receptor expressed on a target cell, forming a heterotrimeric complex.[28] The formation of this complex stimulates the release of a fusogenic peptide, causing the viral membrane to fuse with the membrane of the target host cell.[28] Because binding to CD4 alone can sometimes result in gp120 shedding, gp120 must next bind to co-receptor CCR5 in order for fusion to proceed. The tyrosine-sulfated amino terminus of this co-receptor is the "essential determinant" of binding to the gp120 glycoprotein.[30] The co-receptor also recognizes the V1-V2 region of gp120 and the bridging sheet (an antiparallel, 4-stranded β sheet that connects the inner and outer domains of gp120). The V1-V2 stem can influence "co-receptor usage through its peptide composition as well as by the degree of N-linked glycosylation." Unlike V1-V2 however, the V3 loop is highly variable and thus is the most important determinant of co-receptor specificity.[30] The normal ligands for this receptor, RANTES, MIP-1β, and MIP-1α, are able to suppress HIV-1 infection in vitro. In individuals infected with HIV, CCR5-using viruses are the predominant species isolated during the early stages of viral infection,[31] suggesting that these viruses may have a selective advantage during transmission or the acute phase of disease. Moreover, at least half of all infected individuals harbor only CCR5-using viruses throughout the course of infection.
CCR5 is the primary co-receptor used by gp120 sequentially with CD4. This bind results in gp41, the other protein product of gp160, released from its metastable conformation and inserted into the membrane of the host cell. Although it has not been confirmed, binding of gp120-CCR5 involves two crucial steps: 1) The tyrosine-sulfated amino terminus of this co-receptor is an "essential determinant" of binding to gp120 (as stated previously) 2) Following step 1., there must be reciprocal action (synergy, intercommunication) between gp120 and the CCR5 transmembrane domains.[30]
CCR5 is essential for the spread of the R5-strain of the HIV-1 virus.
Cancer
Expression of CCR5 is induced in breast and prostate epithelial cells upon transformation.[18][19] The induction of CCR5 expression promotes cellular invasion, migration, and metastasis.[5][18][21] The induction of metastasis involves homing to the metastatic site. CCR5 inhibitors including maraviroc and leronlimab have been shown to block lung metastasis of human breast cancer cell lines.[18][36] In preclinical studies of immune competent mice CCR5 inhibitors blocked metastasis to the bones and brain.[19] CCR5 inhibitors also reduce the infiltration of tumor associated macrophages.[37] A Phase 1 clinical study of a CCR5 inhibitor in heavily pretreated patients with metastatic colon cancer demonstrated an objective clinical response and reduction in metastatic tumor burden.[38]
Brain
Increased levels of CCR5 are part of the inflammatory response to stroke and death. Blocking CCR5 with Maraviroc (a drug approved for HIV) may enhance recovery after stroke.[39][40]
In the developing brain, chemokine receptors such as CCR5 influence neuronal migration and connection. After stroke, they seem to decrease the number of connection sites on neurons near the damage.[39]
CCR5-Δ32
CCR5-Δ32 (or CCR5-D32 or CCR5 delta 32) is an allele of CCR5.[41][42]
CCR5 Δ32 is a 32-base-pair deletion that introduces a premature
Recent research indicates that CCR5 Δ32 enhances cognition and memory. In 2016, researchers showed that removing the CCR5 gene from mice significantly improved their memory.[50] CCR5 is a powerful suppressor for neuronal plasticity, learning, and memory; CCR5 over-activation by viral proteins may contribute to HIV-associated cognitive deficits.[51]
Evolutionary history and age of the allele
The CCR5 Δ32 allele is notable for its recent origin, unexpectedly high frequency, and distinct geographic distribution,[52] which together suggest that (a) it arose from a single mutation, and (b) it was historically subject to positive selection.
Two studies have used linkage analysis to estimate the age of the CCR5 Δ32 deletion, assuming that the amount of recombination and mutation observed on genomic regions surrounding the CCR5 Δ32 deletion would be proportional to the age of the deletion.[42][53] Using a sample of 4000 individuals from 38 ethnic populations, Stephens et al. estimated that the CCR5-Δ32 deletion occurred 700 years ago (275-1875, 95% confidence interval). Another group, Libert et al. (1998), used microsatellite mutations to estimate the age of the CCR5 Δ32 mutation to be 2100 years (700-4800, 95% confidence interval). On the basis of observed recombination events, they estimated the age of the mutation to be 2250 years (900-4700, 95% confidence interval).[53] A third hypothesis relies on the north-to-south gradient of allele frequency in Europe, which shows that the highest allele frequency occurred in the Nordic countries and lowest allele frequency in southern Europe. Because the Vikings historically occupied these countries, it may be possible that the allele spread throughout Europe due to the Viking dispersal in the 8th to 10th centuries.[54] Vikings were later replaced by the Varangians in Russia, which may have contributed to the observed east-to-west cline of allele frequency.[52][54]
HIV-1 was initially transmitted from chimpanzees (Pan troglodytes) to humans in the early 1900s in Southeast Cameroon, Africa,
Evidence for a single mutation
Several lines of evidence suggest that the CCR5 Δ32 allele evolved only once.[52] First, CCR5 Δ32 has a relatively high frequency in several different European populations but is comparatively absent in Asian, Middle Eastern and American Indian populations,[42] suggesting that a single mutation occurred after divergence of Europeans from their African ancestor.[42][43][61] Second, genetic linkage analysis indicates that the mutation occurs on a homogeneous genetic background, implying that inheritance of the mutation occurred from a common ancestor.[53] This was demonstrated by showing that the CCR5 Δ32 allele is in strong linkage disequilibrium with highly polymorphic microsatellites. More than 95% of CCR5 Δ32 chromosomes also carried the IRI3.1-0 allele, while 88% carried the IRI3.2 allele. By contrast, the microsatellite markers IRI3.1-0 and IRI3.2-0 were found in only 2 or 1.5% of chromosomes carrying a wild-type CCR5 allele.[53] This evidence of linkage disequilibrium supports the hypothesis that most, if not all, CCR5 Δ32 alleles arose from a single mutational event. Finally, the CCR5 Δ32 allele has a unique geographical distribution indicating a single Northern origin followed by migration. A study measuring allele frequencies in 18 European populations found a North-to-South gradient, with the highest allele frequencies in Finnish and Mordvinian populations (16%), and the lowest in Sardinia (4%).[53]
Positive selection
In the absence of selection, a single mutation would take an estimated 127,500 years to rise to a population frequency of 10%.[42] Estimates based on genetic recombination and mutation rates place the age of the allele between 1000 and 2000 years. This discrepancy is a signature of positive selection.
It is estimated that HIV-1 entered the human population in Africa in the early 1900s,[55] but symptomatic infections were not reported until the 1980s. The HIV-1 epidemic is therefore far too young to be the source of positive selection that drove the frequency of CCR5 Δ32 from zero to 10% in 2000 years.
Protection from bubonic plague. Stephens, et al. (1998), suggest that bubonic plague (Yersinia pestis) had exerted positive selective pressure on CCR5 Δ32.[42] This hypothesis was based on the timing and severity of the Black Death pandemic, which killed 30% of the European population of all ages between 1346 and 1352.[62] After the Black Death, there were less severe, intermittent epidemics. Individual cities experienced high mortality, but overall mortality in Europe was only a few percent.[62][63][64] In 1655-1656 a second pandemic called the "Great Plague" killed 15-20% of London's population.[65][66][dubious ] Importantly, the plague epidemics were intermittent. Bubonic plague is a zoonotic disease, primarily infecting rodents, spread by fleas, and only occasionally infecting humans.[67] Human-to-human infection of bubonic plague does not occur, though it can occur in pneumonic plague, which infects the lungs.[68] Only when the density of rodents is low are infected fleas forced to feed on alternative hosts such as humans, and under these circumstances a human epidemic may occur.[67] Based on population genetic models, Galvani and Slatkin (2003) argue that the intermittent nature of plague epidemics did not generate a sufficiently strong selective force to drive the allele frequency of CCR5 Δ32 to 10% in Europe.[41] To test this hypothesis, Galvani and Slatkin (2003) modeled the historical selection pressures produced by plague and smallpox.[41]
Protection from smallpox. Plague was modeled according to historical accounts,[69][70] while age-specific smallpox mortality was gleaned from the age distribution of smallpox burials in York (England) between 1770 and 1812.[63] Smallpox preferentially infects young, pre-reproductive members of the population since they are the only individuals who are not immunized or dead from past infection. Because smallpox preferentially kills pre-reproductive members of a population, it generates stronger selective pressure than plague.[41] Unlike plague, smallpox does not have an animal reservoir and is only transmitted from human to human.[71][72] The authors calculated that if plague were selecting for CCR5 Δ32, the frequency of the allele would still be less than 1%, while smallpox has exerted a selective force sufficient to reach 10%.
The hypothesis that smallpox exerted positive selection for CCR5 Δ32 is also biologically plausible, since poxviruses, like HIV, enter white blood cells using chemokine receptors.[73] By contrast, Yersinia pestis is a bacterium with a very different biology.
Although Europeans are the only group to have subpopulations with a high frequency of CCR5 Δ32, they are not the only population that has been subject to selection by smallpox, which had a worldwide distribution before it was declared eradicated in 1980. The earliest unmistakable descriptions of smallpox appear in the 5th century A.D. in China, the 7th century A.D. in India and the Mediterranean, and the 10th century A.D. in southwestern Asia.[72] By contrast, the CCR5 Δ32 mutation is found only in European, West Asian, and North African populations.[74] The anomalously high frequency of CCR5 Δ32 in these populations appears to require both a unique origin in Northern Europe and subsequent selection by smallpox.
Potential costs
CCR5 Δ32 can be beneficial to the host in some infections (e.g., HIV-1, possibly smallpox), but detrimental in others (e.g., tick-borne encephalitis, West Nile virus). Whether CCR5 function is helpful or harmful in the context of a given infection depends on a complex interplay between the immune system and the pathogen.[75]
In general, research suggests that the CCR5 Δ32 mutation may play a deleterious role in post-infection inflammatory processes, which can injure tissue and create further pathology.[76] The best evidence for this proposed antagonistic pleiotropy is found in flavivirus infections. In general many viral infections are asymptomatic or produce only mild symptoms in the vast majority of the population. However, certain unlucky individuals experience a particularly destructive clinical course, which is otherwise unexplained but appears to be genetically mediated. Patients homozygous for CCR5 Δ32 were found to be at higher risk for a neuroinvasive form of tick-borne encephalitis (caused by a flavivirus).[77] In addition, functional CCR5 may be required to prevent symptomatic disease after infection with West Nile virus, another flavivirus; CCR5 Δ32 was associated with early symptom development and more pronounced clinical manifestations after infection with West Nile virus.[78]
This finding in humans confirmed a previously observed experiment in an animal model of CCR5 Δ32 homozygosity. After infection with West Nile virus, CCR5 Δ32 mice had markedly increased viral titers in the central nervous system and had increased mortality[79] compared with that of wild-type mice, thus suggesting that CCR5 expression was necessary to mount a strong host defense against West Nile virus.
Medical applications
A genetic approach involving intrabodies that block CCR5 expression has been proposed as a treatment for
This hypothesis was tested in an AIDS patient who had also developed myeloid leukemia, and was treated with chemotherapy to suppress the cancer. A bone marrow transplant containing stem cells from a matched donor was then used to restore the immune system. However, the transplant was performed from a donor with 2 copies of CCR5-Δ32 mutation gene. After 600 days, the patient was healthy and had undetectable levels of HIV in the blood and in examined brain and rectal tissues.[7][81] Before the transplant, low levels of HIV X4, which does not use the CCR5 receptor, were also detected. Following the transplant, however, this type of HIV was not detected either.[7] However, this outcome is consistent with the observation that cells expressing the CCR5-Δ32 variant protein lack both the CCR5 and CXCR4 receptors on their surfaces, thereby conferring resistance to a broad range of HIV variants including HIVX4.[82] After over six years, the patient has maintained the resistance to HIV and has been pronounced cured of the HIV infection.[8]
Enrollment of HIV-positive patients in a clinical trial was started in 2009 in which the patients' cells were genetically modified with a
Inspired by the first person ever to be cured of HIV,
In November 2018,
See also
- Discovery and development of CCR5 receptor antagonists
- Entry inhibitor
- HIV tropism
- Stephen Crohn
- HIV immunity
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000160791 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000079227 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ PMID 31292161.
- PMID 15556703.
- ^ PMID 19213682.
- ^ PMID 21148083.
- PMID 24368413.
- PMID 24597871.
- ^ PMID 24597865.
- ^ a b "CCR5 - chemokine (C-C motif) receptor 5 (gene/pseudogene)". Genetics Home Reference. Archived from the original on 24 September 2009. Retrieved 30 August 2009.
- PMID 8639485.
- ^ PMID 11449371.
- PMID 11734558.
- PMID 14637022.
- PMID 11116158.
- ^ PMID 22637726.
- ^ PMID 25452256.
- S2CID 7976259.
- ^ PMID 29358169.
- PMID 27942440.
- PMID 35154162.
- PMID 9841919.
- PMID 21706028.
- PMID 24623416.
- PMID 18385229.
- ^ S2CID 29364407.
- ^ PMID 19339947.
- ^ a b c d "CCR5 Inhibitors and HIV".
- PMID 17597795.
- ^ PMID 18691983.
- PMID 18691022.
- ^ "Understanding Genetics". The Tech Museum of Innovation, San Jose, CA. 3 December 2021. Archived from the original on 17 June 2021. Retrieved 6 May 2015.
- PMID 18353949.
- PMID 33485378.
- PMID 26238785.
- PMID 27070705.
- ^ a b "HIV drug could improve recovery after stroke". ScienceMag. 21 February 2019. Retrieved 22 February 2019.
- PMID 30794775.
- ^ PMID 14645720.
- ^ PMID 9585595.
- ^ S2CID 21160580.
- S2CID 4370818.
- ^ S2CID 16965624.
- PMID 9388191.
- S2CID 23196048.
- S2CID 24900064.
- ISBN 978-0-8153-4550-3.
- ^ "China's CRISPR twins might have had their brains inadvertently enhanced". Technology Review. Retrieved 22 February 2019.
- PMID 27996938.
- ^ PMID 15715976.
- ^ PMID 9466996.
- ^ PMID 11543895.
- ^ PMID 16728595.
- PMID 10649986.
- ^ Matic S. "HIV/AIDS in Europe: Moving from death sentence to chronic disease management" (PDF). World Health Organization.
- S2CID 32881202.
- PMID 10854218.
- PMID 19472441.
- PMID 9274039.
- ^ PMID 3055286.
- ^ a b Galley C (1998). The Demography of Early Modern Towns: York in the Sixteenth and Seventeenth Centuries. London: MacMillan.
- ^ Hatcher J (1977). Plague, Population, and the English Economy 1348-1530. London: MacMillan.
- ^ "The Great Plague of London, 1665". Contagion, Historical Views of Diseases and Epidemics. Harvard University. Retrieved 2 March 2015.
- ^ "DNA in London Grave May Help Solve Mysteries of the Great Plague". 8 September 2016. Archived from the original on 9 September 2016. Retrieved 18 September 2016.
- ^ PMID 11413636.
- PMID 11614424.
- ^ Russel JS (1948). British Mediaeval Population. Albuquerque: University of New Mexico Press.
- ^ Twigg G (1984). The Black Death: A Biological Reappraisal. London: Batsfod.
- .
- ^ )
- PMID 10583963.
- PMID 16216086.
- PMID 38457490.
- PMID 18179384.
- PMID 18179389.
- PMID 18179388.
- PMID 16230476.
- ^ PMID 10639161.
- ^ Schoofs M (7 November 2008). "A Doctor, a Mutation and a Potential Cure for AIDS". The Wall Street Journal. Retrieved 15 December 2010.
- PMID 14963124.
- ^ "Autologous T-Cells Genetically Modified at the CCR5 Gene by Zinc Finger Nucleases SB-728 for HIV (Zinc-Finger)". U.S. National Institutes of Health. 9 December 2009. Retrieved 30 December 2009.
- ^ Wade N (28 December 2009). "Zinc Fingers Could Be Key to Reviving Gene Therapy". The New York Times. Retrieved 30 December 2009.
- S2CID 34773481.
- PMID 21375684.
- PMID 26251620.
- ^ Begley S (28 November 2018). "Amid uproar, Chinese scientist defends creating gene-edited babies". STAT.
- ^ Collins FS (28 November 2018). "Statement on Claim of First Gene-Edited Babies by Chinese Researcher". National Institutes of Health (NIH). U.S. Department of Health and Human Services.
Further reading
- Wilkinson D (September 1996). "Cofactors provide the entry keys. HIV-1". Current Biology. 6 (9): 1051–1053. S2CID 18710567.
- Broder CC, Dimitrov DS (1996). "HIV and the 7-transmembrane domain receptors". Pathobiology. 64 (4): 171–179. PMID 9031325.
- Choe H, Martin KA, Farzan M, Sodroski J, Gerard NP, Gerard C (June 1998). "Structural interactions between chemokine receptors, gp120 Env and CD4". Seminars in Immunology. 10 (3): 249–257. PMID 9653051.
- Sheppard HW, Celum C, Michael NL, O'Brien S, Dean M, Carrington M, et al. (March 2002). "HIV-1 infection in individuals with the CCR5-Delta32/Delta32 genotype: acquisition of syncytium-inducing virus at seroconversion". Journal of Acquired Immune Deficiency Syndromes. 29 (3): 307–313. PMID 11873082.
- Freedman BD, Liu QH, Del Corno M, Collman RG (2003). "HIV-1 gp120 chemokine receptor-mediated signaling in human macrophages". Immunologic Research. 27 (2–3): 261–276. S2CID 32006625.
- Esté JA (September 2003). "Virus entry as a target for anti-HIV intervention". Current Medicinal Chemistry. 10 (17): 1617–1632. PMID 12871111.
- Gallo SA, Finnegan CM, Viard M, Raviv Y, Dimitrov A, Rawat SS, et al. (July 2003). "The HIV Env-mediated fusion reaction". Biochimica et Biophysica Acta (BBA) - Biomembranes. 1614 (1): 36–50. PMID 12873764.
- Zaitseva M, Peden K, Golding H (July 2003). "HIV coreceptors: role of structure, posttranslational modifications, and internalization in viral-cell fusion and as targets for entry inhibitors". Biochimica et Biophysica Acta (BBA) - Biomembranes. 1614 (1): 51–61. PMID 12873765.
- Lee C, Liu QH, Tomkowicz B, Yi Y, Freedman BD, Collman RG (November 2003). "Macrophage activation through CCR5- and CXCR4-mediated gp120-elicited signaling pathways". Journal of Leukocyte Biology. 74 (5): 676–682. S2CID 11362623.
- Yi Y, Lee C, Liu QH, Freedman BD, Collman RG (2004). "Chemokine receptor utilization and macrophage signaling by human immunodeficiency virus type 1 gp120: Implications for neuropathogenesis". Journal of Neurovirology. 10 (Suppl 1): 91–96. S2CID 9065929.
- Seibert C, Sakmar TP (2004). "Small-molecule antagonists of CCR5 and CXCR4: a promising new class of anti-HIV-1 drugs". Current Pharmaceutical Design. 10 (17): 2041–2062. PMID 15279544.
- Cutler CW, Jotwani R (April 2006). "Oral mucosal expression of HIV-1 receptors, co-receptors, and alpha-defensins: tableau of resistance or susceptibility to HIV infection?". Advances in Dental Research. 19 (1): 49–51. PMID 16672549.
- Ajuebor MN, Carey JA, Swain MG (August 2006). "CCR5 in T cell-mediated liver diseases: what's going on?". Journal of Immunology. 177 (4): 2039–2045. PMID 16887960.
- Lipp M, Müller G (2003). "Shaping up adaptive immunity: the impact of CCR7 and CXCR5 on lymphocyte trafficking". Verhandlungen der Deutschen Gesellschaft für Pathologie. 87: 90–101. PMID 16888899.
- Balistreri CR, Caruso C, Grimaldi MP, Listì F, Vasto S, Orlando V, et al. (April 2007). "CCR5 receptor: biologic and genetic implications in age-related diseases". Annals of the New York Academy of Sciences. 1100 (1): 162–172. S2CID 8437349.
- Madsen HO, Poulsen K, Dahl O, Clark BF, Hjorth JP (March 1990). "Retropseudogenes constitute the major part of the human elongation factor 1 alpha gene family". Nucleic Acids Research. 18 (6): 1513–1516. PMID 2183196.
- Uetsuki T, Naito A, Nagata S, Kaziro Y (April 1989). "Isolation and characterization of the human chromosomal gene for polypeptide chain elongation factor-1 alpha". The Journal of Biological Chemistry. 264 (10): 5791–5798. PMID 2564392.
- Whiteheart SW, Shenbagamurthi P, Chen L, Cotter RJ, Hart GW (August 1989). "Murine elongation factor 1 alpha (EF-1 alpha) is posttranslationally modified by novel amide-linked ethanolamine-phosphoglycerol moieties. Addition of ethanolamine-phosphoglycerol to specific glutamic acid residues on EF-1 alpha". The Journal of Biological Chemistry. 264 (24): 14334–14341. PMID 2569467.
- Ann DK, Wu MM, Huang T, Carlson DM, Wu R (March 1988). "Retinol-regulated gene expression in human tracheobronchial epithelial cells. Enhanced expression of elongation factor EF-1 alpha". The Journal of Biological Chemistry. 263 (8): 3546–3549. PMID 3346208.
- Brands JH, Maassen JA, van Hemert FJ, Amons R, Möller W (February 1986). "The primary structure of the alpha subunit of human elongation factor 1. Structural aspects of guanine-nucleotide-binding sites". European Journal of Biochemistry. 155 (1): 167–171. PMID 3512269.
External links
- Video and text from a PBS documentary about the discovery of CCR5
- "Chemokine Receptors: CCR5". IUPHAR Database of Receptors and Ion Channels. International Union of Basic and Clinical Pharmacology. Archived from the original on 18 January 2021. Retrieved 21 July 2006.
- HIVcoPred A server for prediction of HIV coreceptor usage (CCR5). PLoS ONE 8(4): e61437
- Human CCR5 genome location and CCR5 gene details page in the UCSC Genome Browser.
- Overview of all the structural information available in the PDB for UniProt: P51681 (C-C chemokine receptor type 5) at the PDBe-KB.
