TP63
| View/Edit Human | View/Edit Mouse |
Tumor protein p63, typically referred to as p63, also known as transformation-related protein 63 is a protein that in humans is encoded by the TP63 (also known as the p63) gene.[5][6][7][8]
The TP63 gene was discovered 20 years after the discovery of the p53 tumor suppressor gene and along with p73 constitutes the p53 gene family based on their structural similarity.[9] Despite being discovered significantly later than p53, phylogenetic analysis of p53, p63 and p73, suggest that p63 was the original member of the family from which p53 and p73 evolved.[10]
Function
Tumor protein p63 is a member of the p53 family of transcription factors. p63 -/- mice have several developmental defects which include the lack of limbs and other tissues, such as teeth and mammary glands, which develop as a result of interactions between mesenchyme and epithelium. TP63 encodes for two main isoforms by alternative promoters (TAp63 and ΔNp63). ΔNp63 is involved in multiple functions during skin development and in adult stem/progenitor cell regulation.[11] In contrast, TAp63 has been mostly restricted to its apoptotic function and more recently as the guardian of oocyte integrity.[12] Recently, two new functions have been attributed to TAp63 in heart development[13] and premature aging.[14]
In mice, p63 is required for normal skin development via direct transcription of the membrane protein PERP. TP63 can also regulate PERP expression with TP53 in human cancer.[15]
Oocyte integrity
In oocytes, a unique quality control system has evolved that eliminates by apoptosis those oocytes in which chromosomes do not align correctly, or in which chromosomes cannot be repaired.[16] This monitoring system is conserved from fruit flies and nematodes to humans, and central to this system is the p53 protein family and, in vertebrates in particular, the p63 protein.[16] Oocytes that are unable to repair DNA double-strand breaks produced during meiosis by the process of homologous recombination are eliminated by apoptosis that is linked to p63.[16]
Clinical significance
At least 42 disease-causing mutations in this gene have been discovered.

Both cleft lip with or without a cleft palate and cleft palate only features have been seen to segregate within the same family with a TP63 mutation.[18] Recently, induced pluripotent stem cells have been produced from patients affected by EEC syndromes by cell reprogramming. The defective epithelial commitment could be partially rescued by a small therapeutic compound.[19]
Molecular mechanism
Transcription factor p63 is a key regulator of epidermal keratinocyte proliferation and differentiation. In a recent study, researchers used EEC-patient-derived skin keratinocytes carrying heterozygous p63 DNA-binding domain mutations as the cellular model to characterize the global gene regulatory alteration. The epidermal cell identity was compromised in p63 mutant keratinocytes. Besides, p63-binding loss and loss of active enhancers occurs at a genome-wide scale in patient keratinocytes carrying heterozygous EEC mutations. [20] Besides, using a multi-omics approach, the deregulated function of DNA loops mediated by p63 and CTCF represents an additional layer to the disease mechanism. It seems that a number of loci nearby epidermal genes were organized into a ‘regulatory chromatin hub’ within the chromatin interactions, mediated by CTCF in epidermal keratinocytes. Such hubs contain multiple connecting DNA loops that require not only CTCF binding that is rather static but also binding of cell type-specific TFs, like p63, for the transcriptional activity. In this model, p63 may be essential to make the DNA loops active in transcription. [21]
Vulvar cancer
TP63 has been observed overexpressed in Vulvar Squamous Cell Carcinoma samples, in association with hypermethylation-Induced inactivation of the IRF6 tumor suppressor gene. [22] Indeed, mRNA levels of TP63 tested higher in Vulvar cancer samples when compared with those of normal skin and preneoplastic vulvar lesions, thus underscoring an epigenetic cross-link between IRF6 gene and the oncogene TP63. [22]
Diagnostic utility
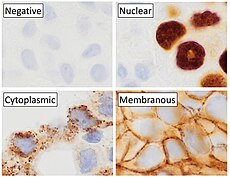
p63
Cytoplasmic staining on immunohistochemistry is seen in cells with muscle differentiation.[26]
Interactions
TP63 has been shown to
Regulation
There is some evidence that the expression of p63 is regulated by the microRNA
See also
- AMACR- another marker for prostate adenocarcinoma
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000073282 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000022510 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- PMID 9774969.
- S2CID 21953916.
- PMID 11181441.
- PMID 11181451.
- PMID 12750249.
- S2CID 10702219.
- PMID 20078223.
- PMID 21335238.
- S2CID 40628077. Archived from the originalon 2014-08-08.
- PMID 19570515.
- S2CID 220631324.
- ^ a b c Gebel J, Tuppi M, Sänger N, Schumacher B, Dötsch V. DNA Damaged Induced Cell Death in Oocytes. Molecules. 2020 Dec 3;25(23):5714. doi: 10.3390/molecules25235714. PMID: 33287328; PMCID: PMC7730327
- PMID 31819097.
- ^ PMID 21331089.
- PMID 23355677.
- PMID 30566872.
- PMID 31164150.
- ^ PMID 27223861.
- PMID 17682568.
- S2CID 9054387.
- PMID 15309021.
- PMID 21623385.)
{{cite journal}}: CS1 maint: multiple names: authors list (link - PMID 12692135.
- PMID 18311128.
- PMID 18848452. Archived from the originalon 2013-04-21.
- S2CID 198263967.
- PMID 32128997.
Further reading
- Little NA, Jochemsen AG (January 2002). "p63". The International Journal of Biochemistry & Cell Biology. 34 (1): 6–9. PMID 11733180.
- van Bokhoven H, McKeon F (March 2002). "Mutations in the p53 homolog p63: allele-specific developmental syndromes in humans". Trends in Molecular Medicine. 8 (3): 133–9. PMID 11879774.
- van Bokhoven H, Brunner HG (July 2002). "Splitting p63". American Journal of Human Genetics. 71 (1): 1–13. PMID 12037717.
- Brunner HG, Hamel BC, van Bokhoven H (October 2002). "P63 gene mutations and human developmental syndromes". American Journal of Medical Genetics. 112 (3): 284–90. PMID 12357472.
- Jacobs WB, Walsh GS, Miller FD (October 2004). "Neuronal survival and p73/p63/p53: a family affair". The Neuroscientist. 10 (5): 443–55. S2CID 39702742.
- Zusman I (2005). "The soluble p51 protein in cancer diagnosis, prevention and therapy". In Vivo. 19 (3): 591–8. PMID 15875781.
- Morasso MI, Radoja N (September 2005). "Dlx genes, p63, and ectodermal dysplasias". Birth Defects Research. Part C, Embryo Today. 75 (3): 163–71. PMID 16187309.
- Barbieri CE, Pietenpol JA (April 2006). "p63 and epithelial biology". Experimental Cell Research. 312 (6): 695–706. PMID 16406339.
- Shalom-Feuerstein R, Lena AM, Zhou H, De La Forest Divonne S, Van Bokhoven H, Candi E, et al. (May 2011). "ΔNp63 is an ectodermal gatekeeper of epidermal morphogenesis". Cell Death and Differentiation. 18 (5): 887–96. PMID 21127502.
External links
- TP73L+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- GeneReviews/NCBI/NIH/UW entry on Ankyloblepharon-Ectodermal Defects-Cleft Lip/Palate Syndrome or AEC Syndrome, Hay-Wells Syndrome. Includes: Rapp–Hodgkin Syndrome
- OMIM entries on AEC
- TP63 human gene location in the UCSC Genome Browser.
- TP63 human gene details in the UCSC Genome Browser.

